Abstract
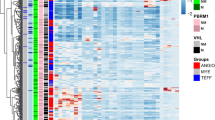
We analysed the expression profiles of 70 kidney tumors of different histological subtypes to determine if these subgroups can be distinguished by their gene expression profiles, and to gain insights into the molecular mechanisms underlying each subtype. In all, 39 clear cell renal cell carcinomas (RCC), seven primary and one metastatic papillary RCC, six granular RCC from old classification, five chromophobe RCC, five sarcomatoid RCC, two oncocytomas, three transitional cell carcinomas (TCC) of the renal pelvis and five Wilms' tumors were compared with noncancerous kidney tissues using microarrays containing 19 968 cDNAs. Based on global gene clustering of 3560 selected cDNAs, we found distinct molecular signatures in clear cell, papillary, chromophobe RCC/oncocytoma, TCC and Wilms' subtypes. The close clustering in each of these subtypes points to different tumorigenic pathways as reflected by their histological characteristics. In the clear cell RCC clustering, two subgroups emerged that correlated with clinical outcomes, confirming the potential use of gene expression signatures as a predictor of survival. In the so-called granular cell RCC (terminology for a subtype that is no longer preferred), none of the six cases clusters together, supporting the current view that they do not represent a single entity. Blinded histological re-evaluation of four cases of ‘granular RCC’ led to their reassignment to other existing histological subtypes, each compatible with our molecular classification. Finally, we found gene sets specific to each subtype. In order to establish the use of some of these genes as novel subtype markers, we selected four genes and performed immunohistochemical analysis on 40 cases of primary kidney tumors. The results were consistent with the gene expression microarray data: glutathione S-transferase α was highly expressed in clear cell RCC, α methylacyl racemase in papillary RCC, carbonic anhydrase II in chromophobe RCC and K19 in TCC. In conclusion, we demonstrated that molecular profiles of kidney cancers closely correlated with their histological subtypes. We have also identified in these subtypes differentially expressed genes that could have important diagnostic and therapeutic implications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Boer JM, Huber WK, Sultmann H, Wilmer F, von Heydebreck A, Haas S, Korn B, Gunawan B, Vente A, Fuzesi L, Vingron M and Poustka A . (2001). Genome Res., 11, 1861–1870.
Brechue WF, Kinne-Saffran E, Kinne RK and Maren TH . (1991). Biochim. Biophys. Acta, 1066, 201–207.
Bugert P and Kovacs G . (1996). Am. J. Pathol., 149, 2081–2088.
Chow WH, Devesa SS, Warren JL and Fraumeni Jr JF . (1999). JAMA, 281, 1628–1631.
Eisen MB and Brown PO . (1999). Methods Enzymol., 303, 179–205.
Eisen MB, Spellman PT, Brown PO and Botstein D . (1998). Proc. Natl. Acad. Sci. USA, 95, 14863–14868.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA, Bloomfield CD and Lander ES . (1999). Science, 286, 531–537.
Grignon DJ, Abdel-Malak M, Mertens WC, Sakr WA and Shepherd RR . (1994). Mod. Pathol., 7, 186–189.
Haddad R, Furge KA, Miller J, Haab BB, Schoumans J, Teh BT, Barr LL and Web CP . (2002). Appl. Genom. Proteom., 1, 51–62.
Hedenfalk I, Duggan D, Chen Y, Radmacher M, Bittner M, Simon R, Meltzer P, Gusterson B, Esteller M, Kallioniemi OP, Wilfond B, Borg A and Trent J . (2001). N. Engl. J. Med., 344, 539–548.
Hegde P, Qi R, Abernathy K, Gay C, Dharap S, Gaspard R, Hughes JE, Snesrud E, Lee and Quackenbush J . (2000). Biotechniques, 29, 548–562.
Jiang Z, Woda BA, Rock KL, Xu Y, Savas L, Khan A, Pihan G, Cai F, Babcook JS, Rathanaswami P, Reed SG, Xu J and Fanger GR . (2001). Am. J. Surg. Pathol., 25, 1397–1404.
Lai LW, Chan DM, Erickson RP, Hsu SJ and Lien YH . (1998). J. Clin. Invest., 101, 1320–1325.
Lewis SE, Erickson RP, Barnett LB, Venta PJ and Tashian RE . (1988). Proc. Natl. Acad. Sci. USA, 85, 1962–1966.
Luo J, Zha S, Gage WR, Dunn TA, Hicks JL, Bennett CJ, Ewing CM, Platz EA, Ferdinandusse S, Wanders RJ, Trent JM, Isaacs WB and De Marzo AM . (2002). Cancer Res., 62, 2220–2226.
Mostfi FK and Davis CJ, (eds) (1998). WHO International Histological Classification of Tumours, Springer: Berlin.
Motzer RJ, Bacik J, Mariani T, Russo P, Mazumdar M and Reuter V . (2002). J. Clin. Oncol., 20, 2376–2381.
Okabe H, Satoh S, Kato T, Kitahara O, Yanagawa R, Yamaoka Y, Tsunoda T, Furukawa and Nakamura Y . (2001). Cancer Res., 61, 2129–2137.
Ono K, Tanaka T, Tsunoda T, Kitahara O, Kihara C, Okamoto A, Ochiai K, Takagi T and Nakamura Y . (2000). Cancer Res., 60, 5007–5011.
Presti Jr JC, Moch H, Reuter VE, Huynh D and Waldman FM . (1996). Genes Chromosomes Cancer, 17, 199–204.
Rhodes DR, Miller JC, Haab BB and Furge KA . (2002). Bioinformatics, 18, 205–206.
Rosenwald A, Wright G, Chan WC, Connors JM, Campo E, Fisher RI, Gascoyne RD, Muller-Hermelink HK, Smeland EB, Giltnane JM, Hurt EM, Zhao H, Averett L, Yang L, Wilson WH, Jaffe ES, Simon R, Klausner RD, Powell J, Duffey PL, Longo DL, Greiner TC, Weisenburger DD, Sanger WG, Dave BJ, Lynch JC, Vose J, Armitage JO, Montserrat E, Lopez-Guillermo A, Grogan TM, Miller TP, LeBlanc M, Ott G, Kvaloy S, Delabie J, Holte H, Krajci P, Stokke T and Staudt LM . (2002). N. Engl. J. Med., 346, 1937–1947.
Rubin MA, Zhou M, Dhanasekaran SM, Varambally S, Barrette TR, Sanda MG, Pienta KJ, Ghosh D and Chinnaiyan AM . (2002). JAMA, 287, 1662–1670.
Sobin LH and Wittekind C, (eds). (2002). TNM Classification of Malignant Tumours, 6th edn. Wiley: New York.
Storkel S, Eble JN, Adlakha K, Amin M, Blute ML, Bostwick DG, Darson M, Delahunt B and Iczkowski K . (1997). Cancer, 80, 987–989.
Takahashi M, Rhodes DR, Furge KA, Kanayama H, Kagawa S, Haab BB and Teh BT . (2001). Proc. Natl. Acad. Sci. USA, 98, 9754–9759.
Van Muijen GN, Warnaar SO and Ponec M . (1987). Exp. Cell Res., 171, 331–345.
Xu J, Stolk JA, Zhang X, Silva SJ, Houghton RL, Matsumura M, Vedvick TS, Leslie KB, Badaro R and Reed SG . (2000). Cancer Res., 60, 1677–1682.
Young AN, Amin MB, Moreno CS, Lim SD, Cohen C, Petros JA, Marshall FF and Neish AS . (2001). Am. J. Pathol., 158, 1639–1651.
Acknowledgements
We thank the Laboratory of DNA and Protein Microarray technology at Van Andel Research Institute (VARI) and Can Gong at the University of Chicago for their technical assistance. We want to acknowledge the contribution of some of the tumors studied by the Cooperative Human Tissue Network (CHTN). We also thank members of the Grand Rapids Urology Study Group including Ken Shockley, John Ludlow, David Kracklau, Philip Wise, Brian Roelof, Jon Curry and pathologists at the Spectrum Health and Metropolitan Hospitals. Finally, we thank Vivve Howell and Rick Hay for kindly reviewing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Takahashi, M., Yang, X., Sugimura, J. et al. Molecular subclassification of kidney tumors and the discovery of new diagnostic markers. Oncogene 22, 6810–6818 (2003). https://doi.org/10.1038/sj.onc.1206869
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206869
Keywords
This article is cited by
-
The 150 most important questions in cancer research and clinical oncology series: questions 57–66
Chinese Journal of Cancer (2017)
-
Molecular and Functional Diagnostic Tools in Precision Oncology for Urological Malignancies
Indian Journal of Surgical Oncology (2017)
-
Glypican 3 overexpression in primary and metastatic Wilms tumors
Virchows Archiv (2015)
-
Reduced cilia frequencies in human renal cell carcinomas versus neighboring parenchymal tissue
Cilia (2013)
-
Functional aspects of primary cilia in signaling, cell cycle and tumorigenesis
Cilia (2013)