Abstract
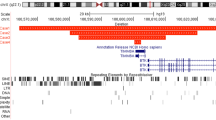
The most frequent genetic alterations described in neuroblastoma (NB) are amplification of MYCN oncogene and deletion of chromosome 1p, although somatic deletions have been demonstrated at other chromosomal intervals. Since loss of heterozygosity (LOH) at distal 4p has been observed in about 20–29% of neuroblastomas, we have evaluated deletions in 41 Italian NB samples by LOH analysis at loci mapping to 4p as follows: pter-D4S2936-D4S412-D4S2957-D4S432-D4S3023-D4S431-cen. Our analysis showed allele losses in eight out of 41 samples (19.5%) and allowed the identification of a smallest region of overlapping deletion (SRO) of 3.0 cM, delimited by D4S412 and D4S3023. Two of these tumors with 4p LOH are from patients belonging to a family with recurrent NB. Interestingly the genotyping of this family revealed an identical haplotype that includes the nonrecombinant loci D4S412, D4S2957 and D4S432 shared by all affected children and demonstrated that this haplotype is retained in the two tumors carrying somatic deletions from patients of this family. Furthermore linkage analysis was performed in two NB families and yielded an overall lod-score of 3.0 in the interval including the haplotype. This provides a confirmatory indication that the region delimited by D4S2936 and D4S3023, which also includes the new defined SRO, may harbor NB predisposing gene/s.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Altura RA, Maris JM, Li H, Boyett JM, Brodeur GM, Look AT . 1997 Genes Chromosomes Cancer 19: 176–184
Bown N, Lastowska M, Cotterill S, O'Neill S, Ellershaw C, Roberts P, Lewis I, Pearson AD . 2001 Med. Pediatr. Oncol. 36: 14–19
Brodeur GM, Seeger RC, Schwab M, Varmus HE, Bishop JM . 1984 Science 224: 1121–1124
Caron H, van Sluis P, Buschman R, Pereira do Tanque R, Maes P, Beks L, de Kraker J, Voute PA, Vergnaud G, Westerveld A, Slater R, Versteeg R . 1996 Hum. Genet. 97: 834–837
Knudson Jr AG . 1971 Proc. Natl. Acad. Sci. USA 68: 820–823
Knudson AG, Strong LC . 1972 Am. J. Hum. Genet. 24: 514–532
Kruglyak L, Daly MJ, Reeve-Daly MP, Lander ES . 1996 Am. J. Hum. Genet. 58: 1347–1363
Lo Cunsolo C, Bicocchi MP, Petti AR, Tonini GP . 2000 Br. J. Cancer 83: 1295–1300
Martinsson T, Sjöberg RM, Hedborg F, Hallstenson K, Nordling M, Kogner P . 1997 Eur. J. Cancer 33: 1997–2001
O'Connell JR, Weeks DE . 1995 Nat. Genet. 11: 402–408
Oude Luttikhuis ME, Powell JE, Rees SA, Genus T, Chughtai S, Ramani P, Mann JR, McConville CM . 2001 Br. J. Cancer 85: 531–537
Perri P, Pession A, Mazzocco M, Scaruffi P, Strigini P, Iolascon A, Albergoni MP, Basso G, Tonini GP . 1997 Eur. J. Cancer 33: 1949–1952
Schwab M, Varmus HE, Bishop JM, Grzeschik KH, Naylor SL, Sakaguchi AY, Brodeur G, Trent J . 1984 Nature 308: 288–291
Schwab M, Praml C, Amler LC . 1996 Genes Chromosomes Cancer 16: 211–229
Srivatsan ES, Ying KL, Seeger RC . 1993 Genes Chromosomes Cancer 7: 32–37
Takita J, Hayashi Y, Kohno T, Shiseki M, Yamaguchi N, Hanada R, Yamamoto K, Yokota J . 1995 Oncogene 11: 1829–1834
Vandesompele J, Van Roy N, Van Gele M, Laureys G, Ambros P, Heimann P, Devalck C, Schuuring E, Brock P, Otten J, Gyselinck J, De Paepe A, Speleman F . 1998 Genes Chromosomes Cancer 23: 141–152
Vandesompele J, Speleman F, Van Roy N, Laureys G, Brinkschmidt C, Christiansen H, Lampert F, Lastowska M, Bown N, Pearson A, Nicholson JC, Ross F, Combaret V, Delattre O, Feuerstein BG, Plantaz D . 2001 Med. Pediatr. Oncol. 36: 5–10
Weiss WA, Aldape K, Mohapatra G, Feuerstein BG, Bishop JM . 1997 EMBO J. 16: 2985–2995
White PS, Maris JM, Beltinger C, Sulman E, Marshall HN, Fujimori M, Kaufman BA, Biegel JA, Allen C, Hillard C, Valentine MB, Look AT, Enomoto H, Sakiyama S, Brodeur GM . 1995 Proc. Natl. Acad. Sci. USA 92: 5520–5524
Acknowledgements
This work was supported by Fondazione Italiana per la Lotta al Neuroblastoma. We are grateful to surgeons, clinicians and pathologists of the Italian Cooperative Group for Neuroblastoma and AIEOP (Associazione Italiana di Ematologia e Oncologia Pediatrica) supplying tumor samples and clinical data, to Dr Katia Mazzocco for MYCN amplification and 1p deletion data and to Dr Roberta Bricchetto for language revision.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Perri, P., Longo, L., Cusano, R. et al. Weak linkage at 4p16 to predisposition for human neuroblastoma. Oncogene 21, 8356–8360 (2002). https://doi.org/10.1038/sj.onc.1206009
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206009
Keywords
This article is cited by
-
Bioinformatics analysis of recurrent deletion regions in neuroblastoma
Medical Oncology (2022)
-
Genetic predisposition and chromosome instability in neuroblastoma
Cancer and Metastasis Reviews (2020)
-
Identification of ALK as a major familial neuroblastoma predisposition gene
Nature (2008)
-
Identification of candidate genes involved in neuroblastoma progression by combining genomic and expression microarrays with survival data
Oncogene (2007)
-
PHOX2B mutations and genetic predisposition to neuroblastoma
Oncogene (2005)