Abstract
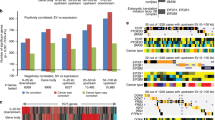
Carcinomas that develop in the pancreatic islets of transgenic mice expressing the SV40 T-antigens (Tag) under transcriptional control of the rat insulin II promoter (RIP) progress through well-characterized stages that are similar to aspects of human tumor progression, including hyperplastic growth, increased angiogenesis and reduced apoptosis1. The latter two stages have been associated with recurrent loss of heterozygosity (LOH)2 and reduced genome copy number3 on chromosomes 9 (LOH9) and 16 (LOH16), aberrations which we believe contribute to these phenotypes. Earlier analyses localized LOH9 to approximately 3 Mb and LOH16 to approximately 30 Mb (both syntenic with human 3q21–q25) but were limited by low throughput and a lack of informative polymorphic markers. Here we show that comparative genomic hybridization to DNA microarrays (array CGH)4,5,6,7 overcomes these limitations by allowing efficient, genome-wide analyses of relative genome copy number. The CGH arrays used in these experiments carried BACs distributed at 2–20-MB intervals across the mouse genome and at higher density in regions of interest. Using array CGH, we further narrowed the loci for LOH9 and LOH16 and defined new or previously unappreciated recurrent regions of copy-number decrease on chromosomes 6, 8 and 14 (syntenic with human chromosomes 12p11–p13, 16q24.3 and 13q11–q32, respectively) and regions of copy-number increase on chromosomes 2 and 4 (syntenic to human chromosomes 20q13.2 and 1p32–p36, respectively). Our analyses of human genome sequences syntenic to these regions suggest that CYP24, PFDN4, STMN1, CDKN1B, PPP2R3 and FSTL1 are candidate oncogenes or tumor-suppressor genes. We also show that irradiation and genetic background influence the spectrum of aberrations present in these tumors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dietrich, W.F. et al. Genome-wide search for loss of heterozygosity in transgenic mouse tumors reveals candidate tumor suppressor genes on chromosomes 9 and 16. Proc. Natl Acad. Sci. USA 91, 9451–9455 (1994).
Parangi, S., Dietrich, W., Christofori, G., Lander, E.S. & Hanahan, D. Tumor suppressor loci on mouse chromosomes 9 and 16 are lost at distinct stages of tumorigenesis in a transgenic model of islet cell carcinoma. Cancer Res. 55, 6071–6076 (1995).
Shi, Y.P. et al. DNA copy number changes associated with characteristic LOH in islet cell carcinomas of transgenic mice. Genes Chromosom. Cancer 19, 104–111 (1997).
Pinkel, D. et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nature Genet. 20, 207–211 (1998).
Pollack, J.R. et al. Genome-wide analysis of DNA copy-number changes using cDNA microarrays. Nature Genet. 23, 41–46 (1999).
Solinas-Toldo, S. et al. Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes Chromosomes Cancer 20, 399–407 (1997).
Heiskanen, M.A. et al. Detection of gene amplification by genomic hybridization to cDNA microarrays. Cancer Res. 60, 799–802 (2000).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Lander, E.S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Venter, J.C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001).
Collins, C. et al. Comprehensive genome sequence analysis of a breast cancer amplicon. Genome Res. 11, 1034–1042 (2001).
Albertson, D.G. et al. Quantitative mapping of amplicon structure by array CGH identifies CYP24 as a candidate oncogene. Nature Genet. 25, 144–146 (2000).
Vainberg, I.E. et al. Prefoldin, a chaperone that delivers unfolded proteins to cytosolic chaperonin. Cell 93, 863–873 (1998).
Iijima, M., Kano, Y., Nohno, T. & Namba, M. Cloning of cDNA with possible transcription factor activity at the G1-S phase transition in human fibroblast cell lines. Acta Med. Okayama 50, 73–77 (1996).
Curmi, P.A. et al. Overexpression of stathmin in breast carcinomas points out to highly proliferative tumours. Br. J. Cancer 82, 142–150 (2000).
Philipp-Staheli, J., Payne, S.R. & Kemp, C.J. p27(Kip1): regulation and function of a haploinsufficient tumor suppressor and its misregulation in cancer. Exp. Cell Res. 264, 148–168 (2001).
Fero, M.L., Randel, E., Gurley, K.E., Roberts, J.M. & Kemp, C.J. The murine gene p27Kip1 is haplo-insufficient for tumour suppression. Nature 396, 177–180 (1998).
Deng, X., Ito, T., Carr, B., Mumby, M. & May, W.S. Jr. Reversible phosphorylation of Bcl2 following interleukin 3 or bryostatin 1 is mediated by direct interaction with protein phosphatase 2A. J. Biol. Chem. 273, 34157–34163 (1998).
Chiang, C.W. et al. Protein phosphatase 2A activates the proapoptotic function of BAD in interleukin-3-dependent lymphoid cells by a mechanism requiring 14-3-3 dissociation. Blood 97, 1289–1297 (2001).
Shibanuma, M., Mashimo, J., Mita, A., Kuroki, T. & Nose, K. Cloning from a mouse osteoblastic cell line of a set of transforming-growth-factor-β 1–regulated genes, one of which seems to encode a follistatin-related polypeptide. Eur. J. Biochem. 217, 13–19 (1993).
Kupprion, C., Motamed, K. & Sage, E.H. SPARC (BM-40, osteonectin) inhibits the mitogenic effect of vascular endothelial growth factor on microvascular endothelial cells. J. Biol. Chem. 273, 29635–29640 (1998).
Efrat, S. et al. Beta-cell lines derived from transgenic mice expressing a hybrid insulin gene-oncogene. Proc. Natl Acad. Sci. USA 85, 9037–9041 (1988).
Han, C.S. et al. Construction of a BAC contig map of chromosome 16q by two-dimensional overgo hybridization. Genome Res. 10, 714–721 (2000).
Shi, Y.P. et al. FISH probes for mouse chromosome identification. Genomics 45, 42–47 (1997).
Korenberg, J.R. et al. Mouse molecular cytogenetic resource: 157 BACs link the chromosomal and genetic maps. Genome Res. 9, 514–523 (1999).
Acknowledgements
This study was supported by USPHS grants CA78601 and R37-CA45234 and by the UCSF Comprehensive Cancer Center Core Grant (CA82103).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hodgson, G., Hager, J., Volik, S. et al. Genome scanning with array CGH delineates regional alterations in mouse islet carcinomas. Nat Genet 29, 459–464 (2001). https://doi.org/10.1038/ng771
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng771
This article is cited by
-
Fstl1/DIP2A/MGMT signaling pathway plays important roles in temozolomide resistance in glioblastoma
Oncogene (2019)
-
Mapping of the chromosomal amplification 1p21-22 in bladder cancer
BMC Research Notes (2014)
-
Genomic copy number alterations in clear cell renal carcinoma: associations with case characteristics and mechanisms of VHL gene inactivation
Oncogenesis (2012)
-
A hidden Markov model-based algorithm for identifying tumour subtype using array CGH data
BMC Genomics (2011)
-
Shared acquired genomic changes in zebrafish and human T-ALL
Oncogene (2011)