Abstract
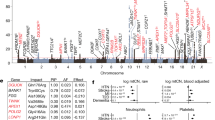
Mammalian mitochondrial DNA (mtDNA) is a high copy-number, maternally inherited genome that codes for a small number of essential proteins involved in oxidative phosphorylation. Mutations in mtDNA are responsible for a broad spectrum of clinical disorders1. The segregation pattern of pathogenic mtDNA mutants is an important determinant of the nature and severity of mitochondrial disease, but it varies with the specific mutation, cell type and nuclear background and generally does not correlate well with mitochondrial dysfunction2,3,4,5,6,7,8,9,10,11. To identify nuclear genes that modify the segregation behavior of mtDNA, we used a heteroplasmic mouse model derived from two inbred strains (BALB/c and NZB; ref. 12), in which we had previously demonstrated tissue-specific and age-dependent directional selection for different mtDNA genotypes in the same mouse13. Here we show that this phenotype segregates in F2 mice from a genetic cross (BALB/c × CAST/Ei) and that it maps to at least three quantitative-trait loci (QTLs). Genome-wide scans showed linkage of the trait to loci on Chromosomes 2, 5 and 6, accounting for 16–35% of the variance in the trait, depending on the tissue and age of the mouse. This is the first genetic evidence for nuclear control of mammalian mtDNA segregation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
DiMauro, S. & Schon, E.A. Mitochondrial DNA mutations in human disease. Am. J. Med. Genet. 106, 18–26 (2001).
Larsson, N.G., Holme, E., Kristiansson, B., Oldfors, A. & Tulinius, M. Progressive increase of the mutated mitochondrial DNA fraction in Kearns–Sayre syndrome. Pediatr. Res. 28, 131–136 (1990).
Boulet, L., Karpati, G. & Shoubridge, E.A. Distribution and threshold expression of the tRNA(Lys) mutation in skeletal muscle of patients with myoclonic epilepsy and ragged-red fibers (MERRF). Am. J. Hum. Genet. 51, 1187–1200 (1992).
Yoneda, M., Chomyn, A., Martinuzzi, A., Hurko, O. & Attardi, G. Marked replicative advantage of human mtDNA carrying a point mutation that causes the MELAS encephalomyopathy. Proc. Natl. Acad. Sci. USA 89, 11164–11168 (1992).
Kawakami, Y. et al. Mitochondrial myopathy with progressive decrease in mitochondrial tRNA(Leu)(UUR) mutant genomes. Ann. Neurol. 35, 370–373 (1994).
Dunbar, D.R., Moonie, P.A., Jacobs, H.T. & Holt, I.J. Different cellular backgrounds confer a marked advantage to either mutant or wild-type mitochondrial genomes. Proc. Natl. Acad. Sci. USA 92, 6562–6566 (1995).
Fu, K. et al. A novel heteroplasmic tRNAleu(CUN) mtDNA point mutation in a sporadic patient with mitochondrial encephalomyopathy segregates rapidly in skeletal muscle and suggests an approach to therapy. Hum. Mol. Genet. 5, 1835–1840 (1996).
Weber, K. et al. A new mtDNA mutation showing accumulation with time and restriction to skeletal muscle. Am. J. Hum. Genet. 60, 373–380 (1997).
Dubeau, F., De Stefano, N., Zifkin, B.G., Arnold, D.L. & Shoubridge, E.A. Oxidative phosphorylation defect in the brains of carriers of the tRNAleu(UUR) A3243G mutation in a MELAS pedigree. Ann. Neurol. 47, 179–185 (2000).
Tang, Y., Manfredi, G., Hirano, M. & Schon, E.A. Maintenance of human rearranged mitochondrial DNAs in long-term cultured transmitochondrial cell lines. Mol. Biol. Cell 11, 2349–2358 (2000).
Rahman, S., Poulton, J., Marchington, D. & Suomalainen, A. Decrease of 3243 A to G mtDNA mutation from blood in MELAS syndrome: a longitudinal study. Am. J. Hum. Genet. 68, 238–240 (2001).
Jenuth, J.P., Peterson, A.C., Fu, K. & Shoubridge, E.A. Random genetic drift in the female germline explains the rapid segregation of mammalian mitochondrial DNA. Nat. Genet. 14, 146–151 (1996).
Jenuth, J.P., Peterson, A.C. & Shoubridge, E.A. Tissue-specific selection for different mtDNA genotypes in heteroplasmic mice. Nat. Genet. 16, 93–95 (1997).
Clayton, D.A. Replication of animal mitochondrial DNA. Cell 28, 693–705 (1982).
Birky, C.W. Jr. The inheritance of genes in mitochondria and chloroplasts: laws, mechanisms, and models. Annu. Rev. Genet. 35, 125–148 (2001).
Chinnery, P.F. et al. Nonrandom tissue distribution of mutant mtDNA. Am. J. Med. Genet. 85, 498–501 (1999).
van den Ouweland, J.M. et al. Mutation in mitochondrial tRNA(Leu)(UUR) gene in a large pedigree with maternally transmitted type II diabetes mellitus and deafness. Nat. Genet. 1, 368–371 (1992).
Petruzzella, V. et al. Extremely high levels of mutant mtDNAs co-localize with cytochrome c oxidase-negative ragged-red fibers in patients harboring a point mutation at nt 3243. Hum. Mol. Genet. 3, 449–454 (1994).
de Stordeur, E., Solignac, M., Monnerot, M. & Mounolou, J.C. The generation of transplasmic Drosophila simulans by cytoplasmic injection: effects of segregation and selection on the perpetuation of mitochondrial DNA heteroplasmy. Mol. Gen. Genet. 220, 127–132 (1989).
Flint, J. & Mott, R. Finding the molecular basis of quantitative traits: successes and pitfalls. Nat. Rev. Genet. 2, 437–445 (2001).
Battersby, B.J. & Shoubridge, E.A. Selection of a mtDNA sequence variant in hepatocytes of heteroplasmic mice is not due to differences in respiratory chain function or efficiency of replication. Hum. Mol. Genet. 10, 2469–2479 (2001).
Meeusen, S. et al. Mgm101p is a novel component of the mitochondrial nucleoid that binds DNA and is required for the repair of oxidatively damaged mitochondrial DNA. J. Cell Biol. 145, 291–304 (1999).
Kaufman, B.A. et al. In organello formaldehyde crosslinking of proteins to mtDNA: identification of bifunctional proteins. Proc. Natl. Acad. Sci. USA 97, 7772–7777 (2000).
Gross, N.J., Getz, G.S. & Rabinowitz, M. Apparent turnover of mitochondrial deoxyribonucleic acid and mitochondrial phospholipids in the tissues of the rat. J. Biol. Chem. 244, 1552–1562 (1969).
Rolink, A.G., Andersson, J. & Melchers, F. Characterization of immature B cells by a novel monoclonal antibody, by turnover and by mitogen reactivity. Eur. J. Immunol. 28, 3738–3748 (1998).
Manly, K.F. & Olson, J.M. Overview of QTL mapping software and introduction to map manager QT. Mamm. Genome. 10, 327–334 (1999).
Basten, C., Weir, B.S. & Zeng, Z.-B. QTL Cartographer: A Reference Manual and Tutorial for QTL Mapping. (Department of Statistics, North Carolina State University, Raleigh, North Carolina, 1997).
Churchill, G.A. & Doerge, R.W. Empirical threshold values for quantitative trait mapping. Genetics. 138, 963–971 (1994).
North, B.V., Curtis, D. & Sham, P.C. A note on the calculation of empirical P values from Monte Carlo procedures. Am. J. Hum. Genet. 71, 439–441 (2002).
Lander, E. & Kruglyak, L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat. Genet. 11, 241–247 (1995).
Acknowledgements
We thank D. Malo, A. Suomalainen, K. Hastings, S. Leary, J. Caron and K. Morgan for discussion and N. Hanna and B. Lauzon for technical assistance. This work was supported by grants from the Canadian Institutes of Health Research and the Howard Hughes Medical Institute to E.A.S. E.A.S. is an international scholar of the Howard Hughes Medical Institute and a Senior Investigator of the Canadian Institutes of Health Research. B.J.B. is supported by a Doctoral Award from the Canadian Institutes of Health Research. J.C.L.-O. is supported by funds to K. Morgan from the Canadian Genetic Diseases Network of Centres of Excellence program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Battersby, B., Loredo-Osti, J. & Shoubridge, E. Nuclear genetic control of mitochondrial DNA segregation. Nat Genet 33, 183–186 (2003). https://doi.org/10.1038/ng1073
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1073
This article is cited by
-
Extreme heterogeneity of human mitochondrial DNA from organelles to populations
Nature Reviews Genetics (2021)
-
Prenatal diagnosis of severe mitochondrial diseases caused by nuclear gene defects: a study in Japan
Scientific Reports (2021)
-
A national perspective on prenatal testing for mitochondrial disease
European Journal of Human Genetics (2014)
-
Manipulation of mtDNA heteroplasmy in all striated muscles of newborn mice by AAV9-mediated delivery of a mitochondria-targeted restriction endonuclease
Gene Therapy (2012)
-
Mitochondrial DNA transcription regulation and nucleoid organization
Journal of Inherited Metabolic Disease (2011)