Abstract
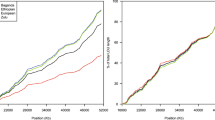
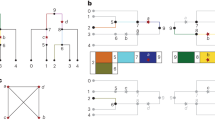
Linkage disequilibrium (LD) analysis is traditionally based on individual genetic markers and often yields an erratic, non-monotonic picture, because the power to detect allelic associations depends on specific properties of each marker, such as frequency and population history. Ideally, LD analysis should be based directly on the underlying haplotype structure of the human genome, but this structure has remained poorly understood. Here we report a high-resolution analysis of the haplotype structure across 500 kilobases on chromosome 5q31 using 103 single-nucleotide polymorphisms (SNPs) in a European-derived population. The results show a picture of discrete haplotype blocks (of tens to hundreds of kilobases), each with limited diversity punctuated by apparent sites of recombination. In addition, we develop an analytical model for LD mapping based on such haplotype blocks. If our observed structure is general (and published data suggest that it may be), it offers a coherent framework for creating a haplotype map of the human genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Rioux, J.D. et al. Hierarchical linkage disequilibrium mapping of a susceptibility gene for Crohn's disease to the cytokine cluster on chromosome 5. Nature Genet. 29, 223–228 (2001).
Templeton, A.R. et al. Recombinational and mutational hotspots within the human lipoprotein lipase gene. Am. J. Hum. Genet. 66, 69–83 (2000).
Jeffreys, A.J., Ritchie, A. & Neumann, R. High resolution analysis of haplotype diversity and meiotic crossover in the human TAP2 recombination hotspot. Hum. Mol. Genet. 9, 725–733 (2000).
Smith, R.A., Ho, P.J., Clegg, J.B., Kidd, J.R. & Thein, S.L. Recombination breakpoints in the human β-globin gene cluster. Blood 92, 4415–4421 (1998).
Jeffreys, A.J., Kauppi, L. & Neumann, R. Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nature Genet. 29, 217–222 (2001).
Sved, J.A. Linkage disequilibrium and homozygosity of chromosome segments in finite populations. Theor. Pop. Biol. 2, 125–141 (1971).
The International SNP Map Working Group. A map of the human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409, 928–933 (2001).
Krawczak, M., Ball, E.V. & Cooper, D.N. Neighboring-nucleotide effects on the rates of germ-line single-base-pair substitution in human genes. Am. J. Hum. Genet. 63, 474–488 (1998).
Nachman, M.W. & Crowell, S.L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000).
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Drysdale, C.M. et al. Complex promoter and coding region β2-adrenergic receptor haplotypes alter receptor expression and predict in vivo responsiveness. Proc. Natl Acad. Sci. USA 97,10483–10488 (2000).
Park, H.Y. et al. Identification of new single–nucleotide polymorphisms in the thrombin receptor gene and their effects on coronary artery diseases in Koreans. Clin. Exp. Pharmacol. Physiol. 27, 690–693 (2000).
Jordanides, N., Eskdale, J., Stuart, R. & Gallagher, G. Allele associations reveal four prominent haplotypes at the human interleukin-6 (IL-6) locus. Genes Immun. 1, 451–455 (2000).
D'Alfonso, S., Rampi, M., Rolando, V., Giordano, M. & Momigliano–Richiardi, P. New polymorphisms in the IL–10 promoter region. Genes Immun. 1, 231–233 (2000).
Bonnen, P.E. et al. Haplotypes at ATM identify coding–sequence variation and indicate a region of extensive linkage disequilibrium. Am. J. Hum. Genet. 67, 1437–1451 (2000).
Moffatt, M.F., Traherne, J.A., Abecasis, G.R. & Cookson, W.O. Single nucleotide polymorphism and linkage disequilibrium within the TCR α/δ locus. Hum. Mol. Genet. 9, 1011–1019 (2000).
Reich, D.R. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Lewontin, R.C. The interaction of selection and linkage. General considerations; heterotic models. Genetics 49, 49–67 (1964).
Hedrick, P.W. Gametic disequilibrium measures: proceed with caution. Genetics 117, 331–341 (1987).
Johnson, G.C.L. et al. Haplotype tagging for the identification of common disease genes. Nature Genet. 29, 233–237 (2001).
Kruglyak, L. & Lander, E.S. High-resolution genetic mapping of complex traits. Am. J. Hum. Genet. 56, 1212–1223 (1995).
Daly, M.J. et al. Genehunter 2.0—a complete linkage analysis system. Am. J. Hum. Genet. 63, A286 (1998).
Dempster, A.P., Laird, N.M. & Rubin, D.B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat. Soc. 39, 1–38 (1977).
Excoffier L. & Slatkin, M. Maximum–likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995).
Clark A.G. Inference of haplotypes from PCR-amplified samples of diploid populations. Mol. Biol. Evol. 7, 111–122 (1990).
Acknowledgements
The authors thank D. Reich, D. Altshuler, J. Hirschhorn, K. Lindblad-Toh and M.P. Reeve for many valuable discussions and comments on the manuscript, A. Kirby for informatics support, the technicians in the inflammatory disease research group at the Whitehead Institute Center for Genome Research for their skilled genotyping work and the anonymous referees for their helpful comments. The authors would also like to thank L. Gaffney for her help in the preparation of this manuscript. This work was supported by research grants from Bristol-Myers Squibb, Millennium Pharmaceutical , and Affymetrix.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Daly, M., Rioux, J., Schaffner, S. et al. High-resolution haplotype structure in the human genome. Nat Genet 29, 229–232 (2001). https://doi.org/10.1038/ng1001-229
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1001-229
This article is cited by
-
Pairwise comparative analysis of six haplotype assembly methods based on users’ experience
BMC Genomic Data (2023)
-
Analysis of protein kinase C (HcPKC) gene expression and single-nucleotide polymorphisms related to inner shell color traits in Hyriopsis cumingii
BMC Genomic Data (2022)
-
Failing the four-gamete test enables exact phasing: the Corners’ Algorithm
Genetics Selection Evolution (2022)
-
Determination of complete chromosomal haplotypes by bulk DNA sequencing
Genome Biology (2021)
-
Polymorphism in the Ras and β-Glucosidase Genes and Their Association with Growth Traits in the Pacific Oyster Crassostrea gigas
Journal of Ocean University of China (2021)