Abstract
The severe mental retardation and speech deficits associated with 22q13 terminal deletions have been attributed in large part to haploinsufficiency of SHANK3, which maps to all 22q13 terminal deletions, although more proximal genes are assumed to have minor effects. We report two children with interstitial deletions of 22q13 and two copies of SHANK3, but clinical features similar to the terminal 22q13 deletion syndrome, including mental retardation and severe speech delay. Both these interstitial deletions are completely contained within the largest terminal deletion, but do not overlap with the nine smallest terminal deletions. These interstitial deletions indicate that haploinsufficiency for 22q13 genes other than SHANK3 can have major effects on cognitive and language development. However, the relatively mild speech problems and normal cognitive abilities of a parent who transmitted her identical interstitial deletion to her more severely affected son suggests that the phenotype associated with this region may be more variable than terminal deletions and therefore contribute to the relative lack of correlation between clinical severity and size of terminal deletions. The phenotypic similarity between the interstitial deletions and non-overlapping small terminal 22q13 deletions emphasizes the general nonspecificity of the clinical picture of the 22q13 deletion syndrome and the importance of molecular analysis for diagnosis.
Similar content being viewed by others
Introduction
Approximately 100 patients have been identified with terminal deletions of 22q13 not involving a ring chromosome.1, 2, 3, 4, 5, 6 Patients with the 22q13 deletion syndrome share common clinical features including severe to profound mental retardation, delay/absence of expressive speech, hypotonia, normal to accelerated growth and minor dysmorphic features (reviewed in Havens et al7). In two studies representing 72 patients, the size of the deletion was determined at the molecular level to range from 130 kb to deletions larger than 9 Mb in size.3, 6 The smallest deletions were found to contain the gene SHANK3, which encodes a synapse structural protein and is located approximately 130 kb from the telomere. Among the 72 patients specifically tested for SHANK3, 61 were hemizyogous at this locus. As all 22q13 deletions, regardless of size, are associated with mental retardation and delay of expressive speech, this implicates SHANK3 as a candidate gene for the major neurological features of this syndrome. This hypothesis is supported by the presence of deletion breakpoints within SHANK3 in the three patients8, 9, 10 and the disruption of SHANK3 in a child with a balanced translocation and all the features of the 22q13 deletion syndrome.11 Recently, dosage imbalance of the SHANK3 gene has been implicated in several patients with autism spectrum disorder.12, 13 Two brothers with a frameshift mutation within SHANK3 showed a severe autistic/mental retardation phenotype that is similar to patients with the 22q13 deletion syndrome, but without any other typical features such as hypotonia.12 A de novo nonsynonymous variant in SHANK3 has also been associated with an autistic phenotype.13 These cases suggest that the SHANK3 gene is sensitive to dosage and that a decrease in dosage can be associated with severe language, cognitive and social defects.
Few correlations have been found between the size of the 22q13 deletion and the clinical features observed, and all patients show mental retardation and severe delay or absence of expressive speech.1, 3, 6 Although one study3 showed the severity of global developmental delay increased with the size of the deletion in 16 patients (plus 17 patients with r(22)), only 4/12 measures of developmental assessment showed some correlation with deletion size in a study of 56 patients.6 If SHANK3 is responsible for most of the neurological abnormalities in these patients, this would imply that deletion of the region proximal to SHANK3 might have a mild phenotype that is largely masked by the terminal deletion of SHANK3.
Almost all of the 22q13 deletions published have been described as terminal. However, there are two cases identified in the literature that have been described as interstitial deletions of 22q13. Romain et al14 cytogenetically identified a patient with an interstitial deletion of 22q13.1–22q13.33 and a phenotype similar to the 22q13 deletion syndrome. The patient shows developmental delay, particularly in motor areas, delay of expressive speech and hypotonia with mild dysmorphic features. The size and the exact location of the deletion in this patient were not characterized at the molecular level, and thus the status of SHANK3 is unknown. In a second patient, a deletion of 22q13.1–22q13.2 was identified by cytogenetics and confirmed by FISH analysis.15 This patient has absence of expressive speech; however, she also has an abnormality of the cochlea and vestibule in both ears that results in bilateral profound hearing loss. She is hypotonic with mild dysmorphic features and developmental delay in motor areas. As the hearing loss in this patient may affect the absence of expressive speech, the analysis of additional cases of interstitial 22q13 deletions is necessary to understand the developmental role of SHANK3 and other genes in this region.
In this report, we identify two patients (A and B) with interstitial 22q13 deletions. Microsatellite, FISH and array-comparative genomic hybridization (array-CGH) analyses were used to determine the size and location of these deletions. Although these interstitial deletions completely overlap with the larger terminal deletions, there is no overlap with many smaller deletions, and SHANK3 is not deleted in either case. Although the patients have a severe phenotype similar to the 22q13 terminal deletion syndrome, the etiology for the neurological features must be due to genes located more proximal to SHANK3. These genes would also be deleted in many patients with 22q13 terminal deletions, and therefore adding to their phenotype. However, the mild speech deficit of the mother of patient B, who has an interstitial deletion identical to her son, highlights the variable effect of the loss of the region proximal to SHANK3 and may explain the lack of correlation of phenotypic severity with the size of the 22q13 deletion.
Materials and methods
Case history
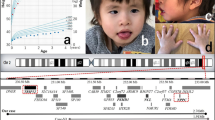
Patient A (Figure 1a and b) is the first child of healthy, non-consanguineous parents. Her features are summarized in Table 1. At 11 months, patient A was referred for neurological evaluation of developmental delay. Severe psychomotor retardation was observed at this time. Examination also identified a slight indication of trigonocephalus. She was a floppy child with a relatively expressionless face. At 4 years and 2 months, she had severe mental retardation with cognitive development corresponding to the sensomotor phase. She was able to grasp, manipulate, throw and explore objects orally. She showed pleasure in sensomotor activities and body contact. She communicates by beating her hands and feet to continue an activity and by hand biting to show discomfort and aggressivity. Her muscle tone and trophism were normal, but she had ligamentous laxity, with an ROT that was normal-vivacious.
Patient B (Figure 1c and d) is the only child of healthy, non-consanguineous Caucasian parents. His phenotype is summarized in Table 1. At three months of age, he was noted to have poor head control, be a poor feeder and had developmental delays. At 8 years of age, patient B attended a special school. He communicated his excitement by flapping his hands. He was able to open a computer by himself and use a mouse. He was able to use a spoon and fork to feed himself and to dress himself (without buttons or zippers).
Both patients A and B have an absence of several features frequently found in the 22q13 deletion syndrome including high pain tolerance, upper respiratory problems and abnormalities of toenails.
The mother of patient B (referred to as BX) was able to attend normal school, but had speech problems and was slow to walk. She had a long face and macrocephaly, but was otherwise normal. The maternal uncle and niece of BX had learning difficulties.
Cytogenetic studies
Peripheral blood lymphocytes were obtained from the probands and relatives for chromosomal analysis. Prometaphase spreads were prepared from synchronized lymphocytes and high-resolution analysis was performed. Metaphase preparations were banded by QFQ, RBA, CBG and Da-DAPI.
Characterization of deletion
Microsatellite loci from Research Genetics were selected throughout the region of the largest previously characterized 22q13 deletion patient (NS from Nesslinger et al16), as well as several more centromeric microsatellite primer pairs. The forward primer was end-labeled with [32P]dATP. PCRs were carried out on patient A and her parental samples using a modification of Research Genetics’ protocol and electrophoresed on either a 6 or 8% polyacrylamide gel. Microsatellites were analyzed by using autoradiographs. Four microsatellite loci from chromosome 15 (D15S822, 15q12, 24.9 Mb; D15S993, 15q22.31, 62 Mb; D15S1020, 15q22.31, 63 Mb; and D15S533, 15q26.2, 95.8 Mb) were analyzed using fluorescently labeled primers and an ABI automated DNA sequencer to test for uniparental disomy of chromosome 15 in patient A.
Cosmid and BAC DNA were used as probes for FISH analysis of metaphase cell preparations of patients A and B as well as BX. The DNA was isolated using QIAGEN® Maxi Kit and was labeled using GIBCOBRL® BioNick Labeling system. The biotinylated probes were detected using the protocol of McTaggart et al,17 with FITC-avidin and anti-avidin purchased from Vector Laboratories.
A customized CGH oligo-array (array no. 10067-4, 4 × 44 K format printed by Agilent, designed by the National Genetics Reference Laboratory in Wessex, UK, http://www.ngrl.org.uk/Wessex/array.htm) was used to further characterize the deletion in patients A, B and EB33. This array maximizes resolution over the whole genome while concentrating on deletion and duplication regions, including 22q13.3 and 15q11.2–q13. The data were analyzed with Agilent's Analytical (v3.3) software platform.
Minisatellite analysis was done for the probe D22S16318 for patient A. Genomic DNA of patient A and both her parents was digested with Sau3A, electrophoresed and transferred to a membrane for Southern blot analysis modified from Sambrook and Russell19 probed with D22S163 labeled with the Ambion® Strip-EZ DNA kit and hybridized in a variation of the Church and Gilbert hybridization solution.20
SHANK3 dosage was also tested by MAPH (Multiplex Amplifiable Probe Hybridisation). MAPH probes were chosen throughout SHANK3, according to the gene structure previously determined,6 using the exon numbering from Durand et al.12 Probes covering 13/24 SHANK3 exons and two 22q11 control probes are given in Table 2. MAPH was performed on 0.5–1 μg genomic DNA using conditions previously described.21 Peak heights were compared by eye for each patient, comparing to a concurrently run normal control and a 22q13 deletion control that was known to have deleted all of SHANK3.
Results
Cytogenetic analysis
Examination of prometaphase spreads of patient A with high-resolution QFQ banding revealed an interstitial deletion at 22q13.1–q13.2 (Figure 2). There was also a supernumerary microchromosome in 85% of the cells. The microchromosome was RBA-negative, C- and Da-DAPI-positive. FISH analysis using the probe D15Z1 showed hybridization signal on the MC, indicating that it originated most likely from chromosome 15, although we have found D15Z1 crosshybridizing polymorphisms on all the other acrocentric chromosomes.22 Uniparental disomy of chromosome 15 was excluded by genotyping the family with four microsatellite loci (D15S822, D15S993, D15S1020 and D15S533). Patient A was heterozygous for all four loci and inherited both a paternal and maternal allele in each case (data not shown).
Examination of the prometaphase spreads of the mother of patient A and of maternal grandmother showed the same supernumerary microchromosome in 79 and 78% of their cells, respectively, whereas their chromosomes 22 were normal. Both of these individuals were phenotypically normal, suggesting that the microchromosome is unlikely to contribute to the phenotype of proband A. The father's karyotype was normal.
Initial analysis of the chromosomes of patient B and BX failed to identify a chromosomal abnormality. When re-examined for subtelomeric abnormalities, the 22q13 interstitial deletion was visibly detected. No abnormalities of the prometaphase spreads of the parents of patient BX were detected indicating that the deletion arose in BX.
Molecular analysis: definition of the deletion
Microsatellite and FISH analyses confirmed that these deletions are interstitial and determined the location of the deletions in comparison to the terminal 22q13 deletion syndrome patients described previously.6 FISH using both cosmids and BACs that span the region of interest was used to characterize the proximal and distal breakpoints of the deletions in patients A, B and BX (Figure 3).
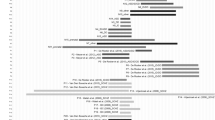
Summary of the size of the A and B interstitial deletions compared to 52 terminal 22q13 deletions described in Wilson et al.6 Four patients from Wilson et al6 were not included due to large ambiguities in the proximal deletion breakpoint (patients 7, 11, 19 and 48). Each patient is represented by a horizontal line. A solid line indicates a confirmed deletion, whereas a dotted line indicates an uninformative region of the deletion. The numbers/letters on the right side of each line indicate individual patients. The thick line at the top represents chromosome 22, with the telomeric region to the right. The microsatellite markers used in this study are labeled by underlined text above the thick line. Cosmids and BACs used in FISH analysis are labeled in bold text, as are oligo-array probes.
Analysis of patient A revealed that microsatellites from D22S274 to D22S270 show inheritance of the paternal allele only, indicating that the deletion is a maternal interstitial deletion. More proximal microsatellites from D22S276 to D22S284 (just proximal to the map in Figure 3) were not deleted, defining its proximal end. Distal microsatellites, from D22S526 to D22S1141, are not deleted confirming that the deletion is interstitial. In patient A, cosmid cN22C6 was present in two copies, defining the telomeric end of the distal breakpoint in this patient. The proximal breakpoint in patient A is flanked by cosmid cN17H6, which is present in only a single copy, and cosmid cN79E12, which is present in two copies. In patients B and BX, BACs 323M22, 388M15, 397C4, 268H5 and 398C22 are all present in a single copy. BAC 29F11, which is the more telomeric BAC used, is present in two copies, verifying that the deletion is indeed interstitial and setting the maximal distal boundary. The proximal breakpoint was identified as being flanked by BAC 323M22, which is deleted, and BAC 85F18, which is not deleted. The deletions in B and BX appeared to be identical.
A customized whole genome CGH oligo-array that concentrates on deletion and duplication regions was used to further narrow down the deletion of patient A and B (Figure 4). The deletion of patient A is located between array clones A_14_P133639 (40.42 Mb) and A_14_P133035 (44.00 Mb) for a maximum deletion size of approximately 3.58 Mb. The array also did not indicate any other abnormalities, including no duplication in the 15q11.2–q13 region or any other acrocentric proximal region, giving further evidence for a lack of euchromatin in the microchromosome of this patient. The overlapping deletion in patient B lies between array clones A_14_P135997 (41.22 Mb) and A_14_P136349 (45.37 Mb) and is up to 4.15 Mb in size.
Characterization of the three deletions using CGH oligo-array. Reduced dosage for probes is shown to the left of the control two copy line, whereas increased dosage is shown to the right. The deleted region is boxed for each patient. Patients A and B show overlapping deletions of 22q13.3 whereas the EB33 deletion is considerably proximal in 22q13.1. Diagrams were produced by the Agilent Analytical (v3.3) software.
Figure 3 shows the location of the interstitial deletions in patients A and B compared to 52 terminal deletions.6 Both of the interstitial deletions are entirely contained within the largest terminal deletion of 22q13 (patient 53) and perhaps others (patients 51, 52, 54 and 55). However, the interstitial deletions of these two patients do not overlap with the nine smallest deletions of the 22q13 deletion syndrome (patients 1, 3, 56, 4, 5, 6, 57, 10 and 8). Table 3 shows a summary of the clinical features of the three interstitial deletion patients (A, B and BX) compared to patients with overlapping and non-overlapping terminal 22q13 deletions.
As deletion of SHANK3 is thought to be responsible for the neurological symptoms of 22q13 terminal deletions, the dosage of SHANK3 in the interstitial deletion patients was specifically tested. Using the minisatellite D22S163, a probe close to the telomere and within SHANK3 (Figure 3, third most distal gene), patient A showed two alleles. The inheritance of two copies of D22S163 in patient A was verified by comparison with the parental alleles. Telomeric microsatellites distal to and including D22S1141 (Figure 3) were also not deleted, confirming that A does not have a terminal deletion. FISH using the Abbot (Vysis) 22q subtelomeric probe D22S163 was performed on chromosomes from patient B and his mother BX. Both B and BX showed two copies of this subtelomeric probe, confirming that SHANK3 is not deleted. MAPH analysis of 13/24 SHANK3 exons further confirmed that this gene was present in two copies in both the patients (Figure 5). Furthermore, the CGH oligo-array showed no indication of a deletion in the SHANK3 region (Figure 4).
An example of MAPH results for SHANK3 exon dosage in a normal control, deletion control and patients A and B. Peaks labeled 6/7, 8, 9, 12, 13, 14, 15/16, 21a, 21b, 21c and 22 represent the dosage of SHANK3 exons. Only in the deletion control individual are these peaks reduced compared to the other control probes from chromosomal regions expected to be present in two copies in all four individuals.
A cell line (EB33) from the patient described in Fujita et al15 was used to compare the interstitial deletion in this patient to the interstitial deletions present in our patients (material was not available for the patient described in Romain et al14). The FISH probes used by Fujita et al (cHKA-19,-44) have not yet been mapped on the published genomic sequence available through NCBI. Using FISH and microsatellite probes to compare the location of this interstitial deletion to that found in our patients, we failed to detect the deletion in EB33 (data not shown). CGH analysis confirmed a more proximal deletion between array clones A_14_P126525 (35.75 Mb) and A_14_P116600 (37.90 Mb) for a maximal size of 2.15 Mb (Figure 4).
Discussion
SHANK3 has been implicated in the neurological features associated with the 22q13 deletion syndrome. SHANK3 (also known as PROSAP2) acts as a scaffold protein in the postsynaptic density of excitatory synapses, linking receptors and signaling proteins to the actin cytoskeleton through a number of protein interactions.23 In most 22q13 deletions, there is relatively little correlation between the severity of cognitive defects and deletion size, as might be expected if the loss of SHANK3 accounts for most neurological features. However, two out of three patients with the smallest deletions8, 10 and the patient with SHANK3 disrupted by a translocation11 have milder cognitive deficits, although speech was severely affected in all. This implies that genes proximal to SHANK3 may make major contributions to the neurological symptoms. To explore this possibility, it is necessary to analyze patients with interstitial 22q13 deletions and two intact copies of SHANK3.
Analysis of patients A and B revealed 22q13 interstitial deletions with distal breakpoints at least 5 Mb proximal to SHANK3, making position effects on SHANK3 unlikely. To rule out a complex rearrangement that could also include SHANK3, the presence of two copies of this gene was confirmed by several methods. Although microsatellite, FISH, MAPH and oligo-array analyses did not test the dosage of all SHANK3 exons, and mutations were not ruled out, it is highly unlikely that these two independent interstitial deletions would also include a small mutation in the distant SHANK3 gene. The distal breakpoints in these patients are more centromeric than the entire deletion in nine patients who show the typical clinical features of the 22q13 deletion syndrome.6 The proximal breakpoints of patients A and B are located in the same region as the breakpoint of the largest 22q13 terminal deletions.6 Thus, both of these interstitial deletions are wholly contained within the largest terminal deletion of patient 53 and perhaps others.
The maximal region of deletion overlap between patients A and B is 2.78 Mb. There are approximately 41 identified or predicted genes in this region (Ensembl release 47). The function and expression pattern of many of these genes are hypothesized or unknown, and there is currently no obvious candidate gene to cause cognitive abnormalities in a dosage-sensitive manner.
The phenotypes of both patients A and B show many similarities to the terminal 22q13 deletion syndrome (Table 3). Like all patients with terminal 22q13 deletion, patients A and B have mental retardation and severe delay or absence of expressive speech even though SHANK3 is not deleted. They both also show hypotonia. Abnormal neurological scans were noted in both A and B, although the findings do not correspond to those identified in any patient with the 22q13 deletion syndrome. Both of these patients have several abnormalities that involve midline structures including abnormalities in the septum pellucidum of both patients, trigonocephaly, multiple nasal and ocular abnormalities, and prominent glabella in patient A. Neither patient has any of the minor clinical features that are commonly associated with both overlapping and non-overlapping terminal 22q13 deletions, such as high pain tolerance and recurrent ear infections, but this could also be explained by nonpenetrance for these features in patients A and B. Comparison of patients A and B with terminal deletions (Table 3) does not reveal an obvious mapping of phenotypes, with the possible exception of macrocephaly to the interstitial region. Overall, the phenotype of patient A and B falls within the typical phenotypic range of 22q13 terminal deletions, despite the fact that the smaller terminal deletions do not overlap with these interstitial deletions, implying that different genes in the more proximal region are responsible.
Two other patients have been reported with interstitial deletions of 22q13. The phenotype of one14 closely matches with that of the terminal 22q13 deletion syndrome with developmental delay, delay of expressive speech, hypotonia, overgrowth, ptosis, full cheeks, as well as several other minor dysmorphisms observed with the 22q13 deletion syndrome. As this case was never characterized molecularly, it is possible that this patient has a terminal rather than interstitial deletion of 22q13. A large terminal deletion studied in Wilson et al6 (patient no. 50, Figure 3) was originally described as an interstitial deletion (data not shown), emphasizing the difficulty of distinguishing the precise location of deletions in the pale 22q13 band. A second interstitial deletion patient originally studied by Fujita et al15 (EB33) has been confirmed to have an interstitial 22q13 deletion by FISH analysis, using cosmids not localized on the published genomic sequence of chromosome 22. Our analysis indicates that the deletion in this patient is more centromeric than the interstitial deletions in patients A and B, and the 56 terminal 22q13 deletions studied in Wilson et al.6
The very mild phenotype observed in patient BX, who has an identical deletion to her more severely affected son patient B, suggests that haploinsufficiency for genes in the interstitial region may be more prone to nonpenetrance and variable expressivity than genes in the SHANK3 region. This is interesting in light of the lack of correlation between the size of the deletion and the severity of the phenotype of patients with the terminal 22q13 deletion syndrome. As patients A, B and BX all have speech deficits of some degree, this suggests that there is a gene in the interstitial region that is important for speech development, and in terminal deletion cases, this gene may variably but significantly add to the effect of haploinsufficiency of SHANK3. The cognitive defects and hypotonia observed in patients A and B, but not in BX, suggests that there are also genes in this region, other than SHANK3, that are responsible for severe neurological symptoms when deleted. These more proximal genes may therefore make a strong contribution to cognitive defects when deleted in many cases of terminal 22q13 deletions, but the severity of that effect may be more variable depending on genetic background, than has been proposed for haploinsufficiency of SHANK3 alone.
The overlap of clinical features between our interstitial patients and the adjacent but non-overlapping small terminal 22q13 deletion cases emphasizes the general nonspecificity of the 22q13 deletion syndrome and the necessity to include molecular analysis in the diagnosis. Non-overlapping deletions of 22q13 can mimic the smaller terminal 22q13 deletions, and although rare, this must be considered when diagnosing patients with developmental delay and delay/absence of expressive speech associated with mild dysmorphic features.
References
Koolen DA, Reardon W, Rosser EM et al: Molecular characterisation of patients with subtelomeric 22q abnormalities using chromosome specific array-based comparative genomic hybridisation. Eur J Hum Genet 2005; 13: 1019–1024.
Lindquist SG, Kirchhoff M, Lundsteen C et al: Further delineation of the 22q13 deletion syndrome. Clin Dysmorphol 2005; 14: 55–60.
Luciani JJ, de Mas P, Depetris D et al: Telomeric 22q13 deletions resulting from rings, simple deletions, and translocations: cytogenetic, molecular, and clinical analyses of 32 new observations. J Med Genet 2003; 40: 690–696.
Manning MA, Cassidy SB, Clericuzio C et al: Terminal 22q deletion syndrome: a newly recognized cause of speech and language disability in the autism spectrum. Pediatrics 2004; 114: 451–457.
Phelan MC, Rogers RC, Saul RA et al: 22q13 deletion syndrome. Am J Med Genet 2001; 101: 91–99.
Wilson HL, Wong ACC, Shaw SR et al: Molecular characterization of the 22q13 deletion syndrome supports the role of haploinsufficiency of SHANK3/PROSAP2 in the major neurological symptoms. J Med Genet 2003; 40: 575–584.
Havens JM, Visootsak J, Phelan MC, Graham Jr JM : 22q13 deletion syndrome: an update and review for the primary pediatrician. Clin pediatr 2004; 43: 43–53.
Anderlid BM, Schoumans J, Anneren G et al: FISH-mapping of a 100-kb terminal 22q13 deletion. Hum Genet 2002; 110: 439–443.
Bonaglia MC, Giorda R, Mani E et al: Identification of a recurrent breakpoint within the SHANK3 gene in the 22q13.3 deletion syndrome. J Med Genet 2006; 43: 822–828.
Wong AC, Ning Y, Flint J et al: Molecular characterization of a 130-kb terminal microdeletion at 22q in a child with mild mental retardation. Am J Hum Genet 1997; 60: 113–120.
Bonaglia MC, Giorda R, Borgatti R et al: Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am J Hum Genet 2001; 69: 261–268.
Durand CM, Betancur C, Boeckers TM et al: Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 2007; 39: 25–27.
Moessner R, Marshall CR, Sutcliffe JS et al: Contribution of SHANK3 mutations to autism spectrum disorder. Am J Hum Genet 2007; 81: 1289–1297.
Romain DR, Goldsmith J, Cairney H, Columbano-Green LM, Smythe RH, Parfitt RG : Partial monosomy for chromosome 22 in a patient with del(22)(pter----q13.1::q13.33----qter). J Med Genet 1990; 27: 588–589.
Fujita Y, Mochizuki D, Mori Y et al: Girl with accelerated growth, hearing loss, inner ear anomalies, delayed myelination of the brain, and del(22)(q13.1q13.2). Am J Med Genet 2000; 92: 195–199.
Nesslinger NJ, Gorski JL, Kurczynski TW et al: Clinical, cytogenetic, and molecular characterization of seven patients with deletions of chromosome 22q13.3. Am J Hum Genet 1994; 54: 464–472.
McTaggart KE, Budarf ML, Driscoll DA, Emanuel BS, Ferreira P, McDermid HE : Cat eye syndrome chromosome breakpoint clustering: identification of two intervals also associated with 22q11 deletion syndrome breakpoints. Cytogenet Cell Genet 1998; 81: 222–228.
Armour JA, Povey S, Jeremiah S, Jeffreys AJ : Systematic cloning of human minisatellites from ordered array charomid libraries. Genomics 1990; 8: 501–512.
Sambrook J, Russell DW : Molecular Cloning: A Laboratory Manual, 3rd edn. Cold Spring Harbor, New York: Cold Spring Harbor Laboratory Press, 2001.
Church GM, Gilbert W : Genomic sequencing. Proc Natl Acad Sci USA 1984; 81: 1991–1995.
Yobb TM, Somerville MJ, Willatt L et al: Microduplication and triplication of 22q11.2: a highly variable syndrome. Am J Hum Genet 2005; 76: 865–876.
Cockwell AE, Jacobs PA, Crolla JA : Distribution of the D15Z1 copy number polymorphism. Eur J Hum Genet 2007; 15: 441–445.
Boeckers TM, Winter C, Smalla KH et al: Proline-rich synapse-associated proteins ProSAP1 and ProSAP2 interact with synaptic proteins of the SAPAP/GKAP family. Biochem Biophys Res Commun 1999b; 264: 247–252.
Acknowledgements
We thank Dr Shuwen Huang and Miss Sarah Beal for the array CGH analysis and Dr N Simon Thomas for the genotyping of chromosome 15. We also wish to thank the families for participating in this research and Professor Yukisige Yanagawa and Dr Yasuko Yamanouchi (Teikyo University School of Medicine) for assistance. This study was supported by a grant from the Canadian Institutes of Health Research (HEM).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wilson, H., Crolla, J., Walker, D. et al. Interstitial 22q13 deletions: genes other than SHANK3 have major effects on cognitive and language development. Eur J Hum Genet 16, 1301–1310 (2008). https://doi.org/10.1038/ejhg.2008.107
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.107
Keywords
This article is cited by
-
Phelan-McDermid syndrome: a classification system after 30 years of experience
Orphanet Journal of Rare Diseases (2022)
-
Investigation of Genetic Polymorphism in Autism Spectrum Disorder: a Pathogenesis of the Neurodevelopmental Disorder
Advances in Neurodevelopmental Disorders (2022)
-
DNA methylation epi-signature is associated with two molecularly and phenotypically distinct clinical subtypes of Phelan-McDermid syndrome
Clinical Epigenetics (2021)
-
Comparison of SHANK3 deficiency in animal models: phenotypes, treatment strategies, and translational implications
Journal of Neurodevelopmental Disorders (2021)
-
Fraternal twins with Phelan-McDermid syndrome not involving the SHANK3 gene: case report and literature review
BMC Medical Genomics (2020)