Abstract
Imprinting is a non-Mendelian form of inheritance where epigenetic modifications control mono-allelic expression depending on the parental origin. Methylation of CpG-dinucleotides at differentially methylated regions (DMRs) is one of the best-studied mechanisms directing expression to one specific parental allele. We studied the methylation patterns of the intergenic (IG)-DMR of DLK1 and GTL2. The methylation marks of the IG-DMR were analysed in human gametes, preimplantation embryos, amniocytes and blood of babies born after intracytoplasmic sperm injection (ICSI) and blood from adults using a bisulphite sequencing technique. In oocytes, the IG-DMR was mainly unmethylated while in sperm cells a generally methylated pattern was detected. This germ-line specific methylation mark was maintained in the preimplantation embryos until the second cleavage stage. Afterwards in the preimplantation embryos, intermediate methylation patterns (26–74% methylation) occurred, which may point to relaxation of the imprints. Intermediate patterns were also present in amniocytes, blood from ICSI babies and adults. We hypothesise that in the early cleavage stage embryo a strict differential methylation pattern is needed for the correct imprint establishment of surrounding imprinted genes. Once correct imprinting of the involved gene(s) is acquired, a more relaxed state of the IG-region is allowed.
Similar content being viewed by others
Introduction
Today nearly 80 imprinted genes have been identified in humans (http://www.geneimprint.com). Imprinted genes, unlike other genes, are characterised by their mono-allelic expression pattern depending on the parental origin of the allele.1 Almost all imprinted genes have differentially methylated regions (DMR) as a mark to distinguish both alleles.2 For primary imprinted genes, the differential methylation patterns arise in gametes, and after fertilisation they are transmitted to form the somatic imprints of the offspring.2, 3 Secondary imprinted genes gain their differentially methylated patterns after fertilisation. The categories of paternally and maternally expressed genes count a comparable number of genes. For most genes from either category the methylation imprint is derived from the oocyte. Only a few imprinted genes with a paternal methylation imprint have been identified. The best-studied paternally methylated region so far is the Igf2-H19 region, which is located on mouse chromosome 7 and is well conserved on human chromosome 11p15.15. Another pair of reciprocally imprinted genes with features that are both unique and in common with the Igf2-H19 locus lies on mouse chromosome 12 and contains the paternally expressed Delta Like homologue 1 (Dlk1) gene and the maternally non-coding RNA-transcript Gene Trap Locus 2 (Gtl2).4, 5, 6, 7, 8, 9, 10 The Dlk1 gene codes for a transmembrane protein that has six epidermal growth factor like repeats in its extracellular domain and shows homology to proteins of the Notch/Delta signalling pathway. Dlk1 plays an important role in normal cellular differentiation and carcinogenesis.11 Gtl2 probably acts as RNA transcript as no consensus sequence for translation was detected.12 Dlk1 and Gtl2 are located within a 1 Mb imprinted cluster containing other imprinted genes such as the paternally expressed Dio3 gene,13 a retrotransposon-like gene (Rtl1),14 several maternally expressed non-coding RNAs, C/D small nucleolar RNAs15 and numerous microRNAs (miRNAs).14, 16
Paulsen et al9 first identified a CpG-rich tandem repeat ‘repeat area 1’ in the intergenic (IG) region of Dlk1/Gtl2 that was conserved between mouse, human and sheep. This repeat area counts seven 24-bp repeats, nine 18-bp repeats and 16 18-bp repeat motifs in the three species, respectively. The IG region was further investigated by Takada et al,10 who identified an 8 kb long IG-DMR associated with a CpG island located 15 kb upstream of Gtl2 and 70 kb downstream of the Dlk1 promotor. This IG-DMR is unmethylated on the maternal allele and hypermethylated on the paternal allele and can be subdivided into three domains of which the last 3′ domain is repeat area 1 which was shown to be differentially methylated. In the mouse, the IG-DMR is a candidate control element for the whole imprinted cluster on chromosome 12. Deletion of the IG-DMR from the maternal chromosome causes bidirectional loss of imprinting of all genes in the cluster including the miRNAs. When inherited from the paternal chromosome, the deletion of the IG-DMR does not affect imprinted gene expression.16, 17
The mechanisms by which methylation imprints are established and maintained in the human genome remain poorly characterised. One of the main blocking factors in this research is the scarcity of human research material, meaning gametes and embryos.
In this work, we studied the methylation profile of the human orthologous repeat area 1 in the IG-region of DLK1-GTL2, located on chromosome 14q32 (Figure 1). Bisulphite sequencing was used to analyse the methylation imprints of the DMR element of this region in gametes, preimplantation embryos, amniocytes and blood of ICSI babies and adults.
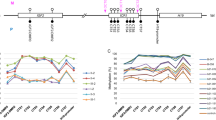
Schematic representation of the analysed IG-DMR on human chromosome 14q32. The vertical bars (D to R) each represent a single CpG site. The primers used in the hemi-nested PCR after bisulphite treatment are shown by grey boxes and the position of the nine repeats is indicated by arrows. The binding of the forward primer IGFIII on the second repeat with one mismatch is indicated with a striped box. M, maternal; P, paternal.
Materials and methods
Samples
The study protocol was approved by the institutional ethical committee. All samples used in the study were donated for research with informed consent of the patients.
Genomic DNA was directly extracted from peripheral blood of adults (control samples for diagnostic tests) and ICSI babies using a QIAmp Blood Maxikit (Qiagen Benelux BL, Venlo, the Netherlands). The isolation of DNA from amniocytes, obtained by routine amniocentesis carried out for prenatal diagnosis at our genetic department (one from an ICSI pregnancy and two from spontaneous pregnancies), was performed using a standard extraction procedure. The blood samples of ICSI babies were obtained in the course of the follow-up studies of ICSI children.
The oocytes, sperm cells and cleavage stage ICSI embryos were obtained and treated as described in Geuns et al.18 The oocytes used were immature at the germinal vesicle (GV) stage or metaphase I (MI) stage, or were left in vitro to spontaneously mature to the metaphase II (MII) stage. Some expanded ICSI blastocysts with a good quality of both inner cell mass (ICM) and trophectoderm were used for laser biopsy of the trophectoderm as described by Cauffman et al 2005.19 Single oocytes or pools of two or three oocytes, single embryos, trophectoderm samples and blanks were transferred to a 1.5 ml tube containing 2 μl alkaline lysis buffer (ALB; 50 mM DTT, 200 mM KOH) and stored at −80°C until use.
Bisulphite treatment
The bisulphite treatment of genomic DNA (blood samples and amniotic cells) was performed according to standard procedures. For the bisulphite treatment of oocytes, sperm cells, embryos and trophectoderm samples a bisulphite sequencing protocol based on low melting point agarose beads was used as described previousls.18
PCR
A hemi-nested PCR protocol was used to amplify the bisulphite converted DNA. In the first round, the forward primer (Eurogentec, Seraing, Belgium) IGFIV (5′-GTGGATTTGTGAGAAATGATTTYGT-3′) (nucleotide position 50975–50999 in GenBank accession number AL117190) labelled with 5′indocarbocyanin (Cy5) was used together with the reverse primer IGRIII (5′-CCATTATAACCAATTACAATACCAC-3′) (51248–51272). The IGFIV primer contained a wobble to allow binding on a methylated as well as on an unmethylated CpG-site. In the second round, the Cy5-labelled forward primer IGFIII (GTTAGTTGTTTGTGGTTTATTAGTTG) (51017–51042) together with the reverse IGRIII primer was used. A PCR-mix containing 0.4 μ M of each primer, 0.2 mM dNTPs (Amersham Pharmacia Biotech, Roosendaal, The Netherlands), 1 × PCR buffer (Applied Biosystems, Nieuwerkerk a/d IJsel, The Netherlands), 2 mM MgCl2 (Applied Biosystems), 1.25 U AmpliTaq DNA Polymerase (Applied Biosystems) in a total volume of 25 μl was used. For, on the one hand genomic DNA and on the other, preimplantation embryos, gametes and trophectoderm cells, the PCR programme in the first round was designed as follows: a denaturation step of 5 min at 94°C followed by 27 and 28 cycles, respectively, of 30 s at 94°C, 30 s at 61.5°C and 30 s at 72°C, and a final extension for 5 min at 72°C. Three microlitres of the first round was used as DNA input for amplification in the second round PCR with the following programme: 5 min denaturation at 94°C followed by 35 cycles of 30 s at 94°C, 30 s at 62°C and 30 s at 72°C, and a final extension step for 5 min at 72°C.
Fragment length analysis of single-cell PCR-products gave two fragments, one of the expected length (256 bp) and another, which is 18 bp or one repeat shorter. For the unmethylated strand, the IGFIII forward primer used in the second round not only anneals (after bisulphite treatment) to the fully orthologous first repeat, but also to the next repeat, despite one mismatch at the 5′ nucleotide of the primer (Figure 1). For the methylated strand, binding at the second repeat would involve two additional mismatches at the methylated CpG-sites D and E and therefore, binding will be much less efficient. In one PCR-cycle, the IGFIII-primers can only bind once to a DNA strand and generate either a full length PCR-fragment or an 18 bp-shorter fragment. No bias will be introduced when considering the totality of short and full-length unmethylated molecules and the totality of (few) short and full-length methylated fragments when calculating the methylation percentage. As further optimisation of the PCR conditions (including testing of other primer sets) was unsuccessful at the single-cell level, both the 256 and 238 bp fragments were cloned; taking into account that the methylation status of CpG-sites D and E could not be determined for the smaller fragment.
The PCR fragments were analysed on an ALFExpress automated sequencer (Amersham Pharmacia Biotech) and positive samples were cloned.
The PCR protocol was validated by the methylation analysis of genomic DNA (adult blood) amplified with a primer set that did not bind to the repeats. Similar results were obtained with this primer set compared to the one we used for our analysis (data not shown).
Cloning and sequencing
The PCR-products were cloned using the TOPO TA cloning kit (Invitrogen, Merelbeke, Belgium). After transformation single colonies were purified on basis of blue/white selection. The length of the insert of 16 colonies was determined by a PCR with M13 primers on lysed bacteria, followed by fragment analysis on a 2% agarose gel. PCR-products of the correct length were then automatically sequenced on the ABI 3100-Avant (Applied Biosystems).
Owing to the bisulphite sequencing procedure and repeats present in the IG-DMR sequence some incomplete or erroneous sequences can be generated. CpG-sites in sequence parts that could not be unequivocally interpreted were indicated as not analysed (NA) in Figure 2.
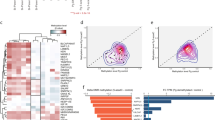
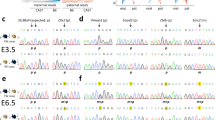
Methylation analysis of the IG-region in (a) oocytes, (b) sperm cells, (c) trophectoderm cells, (d) amniocytes, (e) blood of ICSI-babies and (f) blood of adults. The 15 analysed CpG-dinucleotides are given in the top row of the graph (D–R). The number of clones analysed is mentioned and clones with the same pattern are grouped. For the oocytes and sperm samples, the number of analysed cells in each independent experiment is mentioned in the first column. White boxes are unmethylated CpGs, black boxes are methylated CpGs and NA indicates CpG-sites that could not be analysed. Amniocyte sample 1 and 3 are from spontaneous pregnancies whereas amniocyte sample is from an ICSI pregnancy. GV, germinal vesicle; MI, metaphase I oocyte; MII, metaphase II oocyte; SP, sperm cell.
Checking the intermediate patterns
PCR-products were loaded on a 10% non-denaturating acrylamide gel, and after electrophoresis only bands of the correct length (256 bp) were purified using a standard protocol;20 heteroduplex bands were not cut. Afterwards the purified PCR-product was cloned.
Results
Preliminary tests
Bisulphite sequencing was carried out to analyse the methylation status of the CpG-island in gametes, preimplantation embryos and differentiated cells. The bisulphite sequencing protocol was first optimised for analysis of a small number of cells by embedding the cells in Low Melting Point (LMP) agarose beads. The PCR-protocol was adapted to the single-cell level to allow for analysis of oocytes, sperm cells and preimplantation embryos. With our single-cell PCR-protocol we were able to analyse the methylation status of 15 CpG sites (named D → R) out of 17 CpG sites of the CpG-island. The amplification efficiency of the single-cell protocol, assessed on single-cells, was 17.6% (6/34). Blanks were included in each bisulphite sequencing experiment. In total, 29 blanks were tested and none of them showed amplification. Further optimisation was not successful and this protocol was applied for the methylation analysis of human gametes and embryos, despite its low efficiency.
Gametes
When collecting oocytes, special attention was paid to the complete removal of the zona pellucida and surrounding cumulus cells. Results of the methylation profile of the human IG-DMR in different developmental stages of oogenesis (GV, MI and MII oocytes) were obtained in three independent experiments of single oocytes or pools of two or three oocytes. In the GVs, MI oocytes and MII oocytes a largely unmethylated pattern of 0.0% (0 methylated CpG sites/364 analysed CpG sites), 0.23% (1/433) and 3.87% (12/310) respectively, was found (Figure 2a).
Methylation analysis of four pools of eight or 10 sperm cells showed an overall methylated pattern of 99.3% (421/424) (Figure 2b). Pools with fewer sperm cells did not amplify in our hands, probably because of the more difficult lysis.
Preimplantation embryos
Of the 49 research embryos that were carefully selected (derived from normally fertilised ICSI oocytes and showing normal morphology and developmental timing at the moment of use), 25 cleavage stage embryos (2–10 cells) gave an amplification signal after bisulphite treatment and single-cell PCR. An average methylation pattern of 45.7% (205.8/450.2) was found with 44.4% (114/257) of the clones showing a hypermethylated pattern (74–100%), 49.4% (127/257) a hypomethylated (0–26%) and 6.2% (17/257) an intermediate pattern (26–74%) (Tables 1 and 2). The average methylation percentage was corrected for several factors (see Supplementary Materials) and clones in which four or more CpG-sites out of 15 show a different methylation status were defined as intermediate. Strikingly, the intermediate patterns appeared only after the second cleavage stage (Figure 3). When the results of the methylation analysis of a blastocyst, a compacting and a compacted embryo were pooled, an average methylation pattern of 53.3% (209/392) was found with 37.0% (10/27) of the clones showing a hypermethylated pattern, 25.9% (7/27) showing an unmethylated pattern and 37.0% (10/27) of the clones showing an intermediate pattern. This result was set apart from the mean methylation percentage of the cleavage stage embryos (2–10 cells), because their higher cell number and related higher F-factor (see Supplementary Materials) would bias the average results towards the results of the blastocyst, compacting and compacted embryo.
Schematic overview of the methylation patterns in the 25 cleavage stages embryos, the two morulas and blastocyst. Each bar represents a single embryo and embryos with the same number of blastomeres are grouped. The number under each bar refers to the embryo numbers in Table 1. M, Morula; Bl, Blastocyst.
Trophectoderm cells, amniocytes, genomic DNA isolated from peripheral blood of ICSI babies and adults
As imprinted genes have been reported to play an important role in placental function the selected region was also studied in trophectoderm cells.21, 22, 23 Four trophectoderm samples of good quality blastocysts were separately analysed for their methylation status at the IG-DMR. An average methylation pattern of 27.9% (198/710) was found with 14.3% (7/49) showing a hypermethylated pattern, 61.2% (30/49) of the clones showing a hypomethylated pattern, and 24.5% (12/49) showing an intermediate pattern (Figure 2c, Table 2).
To see whether the loss of the strict differential methylation pattern during the early embryonic period was permanent, further analyses of differentiated cells at different stages of the life cycle (prenatal, postnatal and adult stages) were carried out. In amniocytes samples, an average methylation pattern of 51.6% (222/430) was found, with 41.4% (12/29) of the clones showing a hypermethylated pattern, 44.8% (13/29) a hypomethylated pattern and 13.8% (4/29) an intermediate pattern. Intermediate patterns were seen in all three samples (Figure 2d, Table 2).
The analysed blood samples from ICSI babies, gave an average methylation pattern of the IG-DMR of 48.6% (215/442). Thirty percent (9/30) of the clones showed a hypermethylated pattern, 16.7% (5/30) a hypomethylated pattern and 53.3% (16/30) an intermediate pattern (Figure 2e, Table 2).
The methylation pattern of the IG-DMR in adults was tested on genomic DNA isolated from blood of four healthy adult donors, each analysed in two independent bisulphite sequencing experiment. An average methylation pattern of the IG-DMR of 51.2% (513/1003) was detected with 35.3% (24/68) of the clones showing a hypermethylated pattern, 33.8% (23/68) a hypomethylated pattern and 30.9% (21/68) an intermediate pattern (Figure 2f, Table 2).
Conversion rate
To ensure that the intermediate patterns were not due to incomplete conversion after bisulphite, the conversion rates were determined. The conversion rate for genomic DNA (amniocytes, adult and ICSI blood) was 99.77% (11888 correctly converted cytosines/11915 cytosines). At the single cell level (embryos, oocytes and sperm cells) the conversion rate was 98.81% (25415/25720).
Checking the intermediate patterns
To prove that the intermediate patterns found are a biological phenomenon and not a technical artefact of the bisulphite sequencing technique as described by Sandovici et al,21 further experiments were performed. In a parallel experiment, the PCR-products of bisulphite-treated genomic DNA isolated from blood were either directly used for cloning or were first purified on an acrylamide gel, and only the band of 256 bp was cloned. The acrylamide gel showed a band representing the 238 bp PCR-fragment (18 bp smaller) and other weaker bands, most probably heteroduplexes in addition to the 256 bp fragment. Direct cloning produced three clones out of eight (37.50%) with intermediate patterns. The experiment with the purified 256 bp fragment produced an intermediate pattern for four out of 12 clones (33.33%). All clones in this latter experiment contained the first repeat (CpG sites D and E) as expected for the 256 bp fragment.
Discussion
In this study, we report the methylation status of the IG-DMR of DLK1-GTL2 in human gametes, preimplantation embryos, amniocytes and blood samples (adults and ICSI babies). The development of the single-cell PCR for the IG-DMR was complex, and a rather low amplification efficiency (17.6%) was obtained, compared to similar protocols developed for the SNRPN-gene (28.7%) and the LIT1-gene (25.0%).18, 24 This was most probably due to the numerous tandem repeats present in the sequence of the IG-DMR.
The IG-DMR of DLK1-GTL2 of oocytes at different developmental stages was analysed using the adapted bisulphite sequencing protocol and showed a mainly unmethylated pattern. In spermatozoa on the other hand, a mainly methylated pattern was found. These methylation analysis results indicate that the region under study carries a germ-line mark. The results in human gametes are in agreement with a previous study of the orthologous region in the mouse, in which a germ-line specific methylation mark was also detected, with methylation of the paternal allele.10 After fertilisation, the strict differential methylation status of the IG-DMR was maintained in the embryos until the sixth cell stage. After the second cleavage stage intermediate methylation patterns (26–74%) were found in 6.2% of the cleavage stage embryos (2–10 cells) and in 51.8% of embryos at the morula and blastocyst stage. As imprinted genes have been reported to play an important role in placental functions,22, 23, 25 we also carried out a methylation analysis of the IG-DMR of DLK1-GTL2 in trophectoderm cells biopsied from blastocysts. Here again, part of the clones analysed (28.6%) showed intermediate methylation patterns. This indicates that the intermediate patterns are present both in the embryonic and the extra-embryonic lineage. It is not clear why the average methylation percentage in trophectoderm cells was lower than in embryos and blood samples. Further analyses of extra-embryonic tissues at different developmental stages are required to clarify this.
In mouse, the occurrence of intermediate methylation patterns for this region has not been previously reported. Analysis of mouse embryos (embryonic day 13.5) with maternal and paternal uniparental disomy (UPD) for chromosome 12 revealed strictly unmethylated and methylated patterns, respectively.10
Bisulphite sequencing of the IG-DMR of DLK1-GTL2 at later developmental stages (genomic DNA isolated from amniocytes, blood of ICSI-babies or from adults) also produced clones with intermediate methylation patterns. The percentage of intermediate patterns varied among the samples; this most probably only reflects the limitations of the methodology. It was not possible to determine the parental origin of the intermediate patterns. Although there was one SNP (RS967189, C/G polymorphism) available in the studied region, only the G allele was observed in our samples. In humans, there are three papers describing the methylation status of repeat area 1 of the IG-DMR. The publication by Lin et al,17 investigated the methylation status of a flanking region (including CpG-site T) using methylation-sensitive restriction enzymes and a Southern blot probe encompassing our selected region. The authors' analysis of DNA from mUPD14 and pUPD14 patients showed a paternal methylation mark, but intermediate patterns were not reported. These results were similar to the data obtained from mouse studies.10 The second paper involved the methylation analysis of CpG-sites D to K in four normal control blood samples using the bisulphite sequencing technique. Here, an overall methylation percentage of 32% was found; however, the presentation of the data did not allow checking individual clones for intermediate patterns.26 The third paper examined the methylation status of the IG-DMR (CpG site D-R) in four peripheral blood samples using the bisulphite sequencing technique. Detailed information about one representative sample is given and showed intermediate patterns in three of the eight clones analysed.27
The possibility exists that the intermediate patterns represent bacterial recombination artefacts. The amplification of bisulphite-treated DNA can generate heteroduplexes, as the parental template strands only differ at specific CpG-sites. Upon cloning, the heteroduplexes can be converted into a single hybrid sequence by the host's mismatch repair mechanism.21 We were able to exclude this possibility by demonstrating that cloning of purified PCR-samples as well as non-purified PCR samples produced a similar percentage of intermediate methylation patterns. It was also ruled out that the intermediate patterns stemmed from incomplete bisulphite conversion.
Other reports have also shown that the maintenance of paternal and maternal methylation patterns was not absolute. Studies at the imprint control region of H19/Igf2 in mice as well as in the human showed around 5 and 6% intermediate methylation patterns (40–50%) respectively.28, 29 These percentages are lower than the relaxation of allele-specific methylation observed at the IG-DMR.
Relaxation of imprinting, indicating biallelic expression of a gene that is normally expressed from one parental allele, and associated altered DNA methylation patterns at DMRs has been described as a polymorphic trait among individuals, being tissue specific or age related.27, 30, 31, 32, 33 Our data rather indicate a common phenomenon as intermediate patterns of the IG-DMR have been observed in different tissue samples and in all the different subjects analysed. Relaxation of the IG-DMR methylation patterns may not necessarily imply relaxation of imprinting expression at the cluster since other epigenetic modifications can contribute to the mono-allelic expression of these imprinted genes. We hypothesise that a differential methylation pattern is needed in the early cleavage stage embryo for the correct imprint establishment of (an) other imprinted gene(s) in the DLK1-GTL2 locus. Once correct imprinting of the involved genes is acquired, the IG-DMR may become redundant and a more relaxed state may be allowed.
Several studies have demonstrated that in vitro culture systems and embryo manipulations have a detrimental influence on imprinting mechanisms in animal models.34, 35, 36 The influence of in vitro culture conditions and the ICSI-procedure on the methylation patterns of the oocytes and embryos in our experiments cannot be excluded. However, in humans these embryos are the only source of research material that can be used. The proportion of intermediate patterns in the preimplantation embryos increases with time in culture. Similarly, analysis of somatic cells of pre- and postnatal stages revealed intermediate methylation patterns similar to those in the preimplantation embryos, which led us to believe that these patterns indeed arise in the preimplantation embryo physiologically, and are not an artefact of the in vitro manipulations. Recent publications about a higher incidence of BWS and AS, both imprinting disorders with a low incidence, after ICSI and IVF37, 38, 39, 40, 41, 42, 43, 44, 45, 46 address the risks of ART and highlight the need for basic molecular studies of the imprinting processes in the human.
Accession codes
References
Constancia M, Hemberger M, Hughes J et al: Placental-specific IGF-II is a major modulator of placental and fetal growth. Nature 2002; 417: 945–948.
Reik W, Santos F, Dean W : Mammalian epigenomics: reprogramming the genome for development and therapy. Theriogenology 2003; 59: 21–32.
Reik W, Dean W, Walter J : Epigenetic reprogramming in mammalian development. Science 2001; 293: 1089–1093.
Kobayashi S, Wagatsuma H, Ono R et al: Mouse Peg9/Dlk1 and human PEG9/DLK1 are paternally expressed imprinted genes closely located to the maternally expressed imprinted genes: mouse Meg3/Gtl2 and human MEG3. Genes Cells 2000; 5: 1029–1037.
Schmidt JV, Matteson PG, Jones BK, Guan XJ, Tilghman SM : The Dlk1 and Gtl2 genes are linked and reciprocally imprinted. Genes Dev 2000; 14: 1997–2002.
Wylie AA, Murphy SK, Orton TC, Jirtle RL : Novel imprinted DLK1/GTL2 domain on human chromosome 14 contains motifs that mimic those implicated in IGF2/H19 regulation. Genome Res 2000; 10: 1711–1718.
Takada S, Tevendale M, Baker J et al: Delta-like and gtl2 are reciprocally expressed, differentially methylated linked imprinted genes on mouse chromosome 12. Curr Biol 2000; 10: 1135–1138.
Charlier C, Segers K, Wagenaar D et al: Human-ovine comparative sequencing of a 250-kb imprinted domain encompassing the callipyge (clpg) locus and identification of six imprinted transcripts: DLK1, DAT, GTL2, PEG11, antiPEG11, and MEG8. Genome Res 2001; 5: 850–862.
Paulsen M, Takada S, Youngson NA et al: Comparative sequence analysis of the imprinted Dlk1-Gtl2 locus in three mammalian species reveals highly conserved genomic elements and refines comparison with the Igf2-H19 region. Genome Res 2001; 11: 2085–2094.
Takada S, Paulsen M, Tevendale M et al: Epigenetic analysis of the Dlk1-Gtl2 imprinted domain on mouse chromosome 12: implications for imprinting control from comparison with Igf2-H19. Hum Mol Genet 2002; 11: 77–86.
Laborda J : The role of the epidermal growth factor-like protein dlk in cell differentiation. Histol Histopathol 2000; 15: 119–129.
Schuster-Gossler K, Bilinski P, Sado T, Furguson-Smith A, Gossler A : The mouse Gtl2 gene is differentially expressed during embryonic development, encodes multiple alternatively spliced transcripts, and may act as an RNA. Dev Dyn 1998; 212: 214–228.
Tsai CE, Lin SP, Ito M, Takagi N, Takada S, Ferguson-Smith AC : Genomic imprinting contributes to thyroid hormone metabolism in the mouse embryo. Curr Biol 2002; 12: 1221–1226.
Seitz H, Youngson N, Lin SP et al: Imprinted microRNA genes transcribed antisense to a reciprocally imprinted retrotransposon-like gene. Nat Genet 2003; 34: 261–262.
Cavaille J, Seitz H, Paulsen M, Ferguson-Smith AC, Bachellerie JP : Identification of tandemly-repeated C/D snoRNA genes at the imprinted human 14q32 domain reminiscent of those at the Prader-Willi/Angelman syndrome region. Hum Mol Genet 2002; 11: 1527–1538.
Seitz H, Royo H, Bortolin ML, Lin SP, Ferguson-Smith AC, Cavaille J : A large imprinted microRNA gene cluster at the mouse Dlk1-Gtl2 domain. Genome Res 2004; 14: 1741–1748.
Lin SP, Youngson N, Takada S et al: Asymmetric regulation of imprinting on the maternal and paternal chromosomes at the Dlk1-Gtl2 imprinted cluster on mouse chromosome 12. Nat Genet 2003; 35: 97–102.
Geuns E, De Rycke M, Van Steirteghem A, Liebaers I : Methylation imprints of the imprint control region of the SNRPN-gene in human gametes and preimplantation embryos. Hum Mol Genet 2003; 12: 2873–2879.
Cauffman G, Van de Velde H, Liebaers I, Van Steirteghem A : DAZL expression in human oocytes, preimplantation embryos and embryonic stem cells. Mol Hum Reprod 2005; 11: 405–411.
Maxam A, Gilbert W : Sequencing end-labeled DNA with base specific chemical cleavages. Methods Enzymol 1980; 65: 499–559.
Sandovici I, Leppert M, Hawk PR, Suarez A, Linares Y, Sapienza C : Familial aggregation of abnormal methylation of parental alleles at the IGF2/H19 and IGF2R differentially methylated regions. Hum Mol Genet 2003; 12: 1569–1578.
Tycko B : Imprinted genes in placental growth and obstetric disorders. Cytogenet Genome Res 2006; 113: 271–278.
Smith FM, Garfield AS, Ward A : Regulation of growth and metabolism by imprinted genes. Cytogenet Genome Res 2006; 113: 279–291.
Geuns E, Hilven P, Van Steirteghem A, Liebaers I, De Rycke M : Methylation analysis of KvDMR1 in human oocytes. J Med Genet 2006, E-pub ahead of print, 1 September 2006.
Reik W, Constancia M, Fowden A et al: Regulation of supply and demand for maternal nutrients in mammals by imprinted genes. J Physiol 2003; 547: 35–44.
Astuti D, Latif F, Wagner K et al: Epigenetic alteration at the DLK1-GTL2 imprinted domain in human neoplasia: analysis of neuroblastoma, phaeochromocytoma and Wilms' tumour. Br J Cancer 2005; 92: 1574–1580.
Kawakami T, Chano T, Minami K, Okabe H, Okada Y, Okamoto K : Imprinted DLK1 is a putative tumor suppressor gene and inactivated by epimutation at the region upstream of GTL2 in human renal cell carcinoma. Hum Mol Genet 2006; 15: 821–830.
Bowman AB, Levorse JM, Ingram RS, Tilghman SM : Functional characterization of a testis-specific DNA binding activity at the H19/Igf2 imprinting control region. Mol Cell Biol 2003; 23: 8345–8351.
Kerjean A, Dupont JM, Vasseur C et al: Establishment of the paternal methylation imprint of the human H19 and MEST/PEG1 genes during spermatogenesis. Hum Mol Genet 2000; 9: 2183–2187.
Xu Y, Goodyer CG, Deal C, Polychronakos C : Functional polymorphism in the parental imprinting of the human IGF2R gene. Biochem Biophys Res Commun 1993; 15: 747–754.
Jinno Y, Yun K, Nishiwaki K et al: Mosaic and polymorphic imprinting of the WT1 gene in humans. Nat Genet 1994; 6: 305–309.
Weber M, Milligan L, Delalbre A et al: Extensive tissue-specific variation of allelic methylation in the Igf2 gene during mouse fetal development: relation to expression and imprinting. Mech Dev 2001; 101: 133–141.
Sakatani T, Wei M, Katoh M et al: Epigenetic heterogeneity at imprinted loci in normal populations. Biochem Biophys Res Commun 2001; 283: 1124–1130.
Young LE, Fernandes K, McEvoy TG et al: Epigenetic change in IGF2R is associated with fetal overgrowth after sheep embryo culture. Nat Genet 2001; 27: 153–154.
Doherty AS, Mann MR, Tremblay KD, Bartolomei MS, Schultz RM : Differential effects of culture on imprinted H19 expression in the preimplantation mouse embryo. Biol Reprod 2000; 62: 1526–1535.
Khosla S, Dean W, Brown D, Reik W, Feil R : Culture of preimplantation mouse embryos affects fetal development and the expression of imprinted genes. Biol Reprod 2001; 64: 918–926.
Cox GF, Bürger J, Lip V et al: Intracytoplasmic sperm injection may increase the risk of imprinting defects. Am J Hum Genet 2002; 71: 162–164.
De Rycke M, Liebaers I, Van Steirteghem : A epigenetic risks related to assisted reproductive technologies – Risk analysis and epigenetic inheritance. Hum Reprod 2002; 17: 2487–2494.
DeBaun MR, Niemitz EL, Feinberg AP : Association of in vitro fertilization with Beckwith–Wiedemann syndrome and epigenetic alterations of LIT1 and H19. Am J Hum Genet 2003; 72: 156–160.
Gicquel C, Gaston V, Mandelbaum J, Siffroi JP, Flahault A, Le Bouc Y : In vitro fertilization may increase the risk of Beckwith–Wiedemann syndrome related to the abnormal imprinting of the KCN1OT gene. Am J Hum Genet 2003; 72: 1338–1341.
Maher ER, Brueton LA, Bowdin SC et al: Beckwith–Wiedemann syndrome and assisted reproduction technology (ART). J Med Genet 2003; 40: 62–64.
Orstavik KH, Eiklid K, Van der Hagen Cb et al: Another case of imprinting defect in a girl with Angelman syndrome who was conceived by intracytoplasmic semen injection. Am J Hum Genet 2003; 72: 218–219.
Halliday J, Oke K, Breheny S, Algar E, Amor D : Beckwith–Wiedemann syndrome and IVF: a case–control study. Am J Hum Genet 2004; 75: 526–528.
Chang AS, Moley KH, Wangler M, Feinberg AP, Debaun MR : Association between Beckwith–Wiedemann syndrome and assisted reproductive technology: a case series of 19 patients. Fert Steril 2005; 83: 349–354.
Ludwig M, Katalinic A, Gross S, Sutcliffe A, Varon R, Horsthemke B : Increased prevalence of imprinting defects in patients with Angelman syndrome born to subfertile couples. J Med Genet 2005; 42: 289–291.
Sutcliffe AG, Peters CJ, Bowdin S et al: Assisted reproductive therapies and imprinting disorders – a preliminary British survey. Hum Reprod 2006; 21: 1009–1011.
Acknowledgements
We thank the staff at the IVF-lab and Centre for Medical Genetics for providing the research material that was donated by the patients. We also thank Professor Dr K Sermon and Dr W Lissens for helpful discussions and critically reading this manuscript. Furthermore, we are grateful to M Whitburn of the Language Education Centre at our University for proofreading the text of this article. Our work was supported by the Fund for Scientific Research, Flanders, Belgium, and the Scientific Council of the Vrije Universiteit Brussel.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Geuns, E., De Temmerman, N., Hilven, P. et al. Methylation analysis of the intergenic differentially methylated region of DLK1-GTL2 in human. Eur J Hum Genet 15, 352–361 (2007). https://doi.org/10.1038/sj.ejhg.5201759
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201759
Keywords
This article is cited by
-
Effects of paternal exposure to cigarette smoke on sperm DNA methylation and long-term metabolic syndrome in offspring
Epigenetics & Chromatin (2022)
-
Analysis of DNA methylation in single circulating tumor cells
Oncogene (2017)
-
New insights into the imprinted MEG8-DMR in 14q32 and clinical and molecular description of novel patients with Temple syndrome
European Journal of Human Genetics (2017)
-
High Frequency of Imprinted Methylation Errors in Human Preimplantation Embryos
Scientific Reports (2015)
-
The specification of imprints in mammals
Heredity (2014)