Abstract
It is well known that a variety of genetic changes influence the development and progression of cancer. These changes may result from inherited or spontaneous mutations that are not corrected by repair mechanisms prior to DNA replication. It is increasingly clear that so called epigenetic effects that do not affect the primary sequence of the genome also play an important role in tumorigenesis. This was supported initially by observations that cancer genomes undergo changes in their methylation state and that control of parental allele-specific methylation and expression of imprinted loci is lost in several cancers. Many loci acquiring aberrant methylation in cancers have since been identified and shown to be silenced by DNA methylation. In many cases, this mechanism of silencing inactivates tumour suppressors as effectively as frank mutation and is one of the cancer-predisposing hits described in Knudson's two hit hypothesis. In contrast to mutations which are essentially irreversible, methylation changes are reversible, raising the possibility of developing therapeutics based on restoring the normal methylation state to cancer-associated genes. Development of such therapeutics will require identifying loci undergoing methylation changes in cancer, understanding how their methylation influences tumorigenesis and identifying the mechanisms regulating the methylation state of the genome. The purpose of this review is to summarise what is known about these issues.
Similar content being viewed by others
I. Regulation of DNA methylation in normal development
Introduction
In the mammalian genome, cytosine residues may be methylated at the 5′ carbon. This occurs most commonly at 5′-CG-3′ dinucleotides but cytosine methylation can be found at 5′-CA-3′ or 5′-CT-3′ residues.1 Since 5′-CG-3 is palindromic, the cytosine on one or both DNA strands may be methylated (hemi-methylated and homo-methylated respectively). DNA methylation is an epigenetic modification that does not alter the primary sequence of DNA and is critical for normal development, gene expression patterns and genomic stability (see Table 1). DNA methylation patterns are dynamic – unmethylated sequences can become methylated and methyl groups can be lost. For example, differences in methylation patterns have been described between oocytes and generally more highly methylated sperm DNA; post-fertilisation development is characterised by waves of genome-wide demethylation in early stages and subsequent remethylation before implantation; CpGs within promoters of many genes on the inactive X-chromosome in female cells are methylated and may be part of the X-inactivation mechanism; differentially methylated regions (DMRs) have been described for imprinted genes where the allele from one parent is methylated but the other is not (see below for more details). Of particular relevance to this review is the fact that perturbations in DNA methylation have been associated with abnormal developmental processes including cancer.
Establishment of methylation patterns(DNMTs and their actions)
Methylation is established by de novo methyltransferases (see Figure 1). Replication of homomethylated DNA produces hemimethylated DNA in which one strand of the DNA remains methylated and the newly synthesised is unmethylated. Hemimethylated DNA can become homomethylated by maintenance methyltransferases which place a methyl group at the 5′-CG-3 complimentary to a methylated 5′-CG-3. The first DNA methyltransferase identified, DNMT1, was shown to have both de novo methylation2 as well as maintenance activity, although the de novo activity is much weaker than the maintenance activity. In cells lacking DNMT1 maintenance methylation is not abolished indicating other DNMTs have maintenance activity.3 Two more potent de novo methyltransferases have been identified: DNMT3A and DNMT3B. Neither is required for maintenance activity since ES cells deficient for either DNMT maintain the pre-existing methylation patterns.4 The importance of each of these enzymes for normal development has been demonstrated in mouse mutants that lack the individual genes. Mouse embryos homozygous for mutant alleles of Dnmt1, Dnmt3a or Dnmt3b die early in development either before birth (Dnmt1, Dnmt3b), or a few weeks after birth (Dnmt3a). These experiments dramatically revealed the importance of DNA methylation in normal development, but also identified individual target sequences for each of the enzymes. For example, DNMT3B is required for the methylation of centromeric minor satellite repeat sequences and both DNMT3A and DNMT3B are involved in methylation of C-type retroviral repeats, IAP repeats, major satellite DNA and the differentially methylated region in Igf2. Sequence specificity of DNMT3A and DNMT3B seen in in vivo experiments was not seen in in vitro assays indicating the requirement for additional factors that direct methyltransferase activity.5 Proteins with homology to the other DNMTs (DNMT2 and DNMT3L) and alternatively spliced forms of DNMT1 and DNMT3B with altered function have been reported,6,7 however their functions are not completely understood.
DNA demethylation may occur actively by an enzyme with demethylating activity, or passively by several rounds of replication in the absence of maintenance methyltransferase activity. There is evidence that both processes occur. Active demethylation occurs predominantly on the paternally transmitted chromosomes in zygotes while maternal chromosomes undergo passive demethylation in later cleavage stages.8
Trans-acting regulatory factors
Identifying the factors that regulate establishment and maintenance of DNA methylation in both normal and cancer genomes will be important for identifying targets of therapeutic intervention. Few details are known about trans-acting factors regulating methylation in mammals and other organisms. While the DNMTs are clearly central to methylation establishment and maintenance, additional trans-acting factors regulating the timing and targets of their activity must exist, otherwise, methylation patterns would be static and homogenous across the genome. In Neurospora crassa, a histone methyltransfease is required for DNA methylation.9 The observations that DNMT1 can be detected in complexes with Rb, E2F1 and HDAC110 and also with MBD2 and MBD3 localised to replication forks in late S phase11 highlight these associated factors as potential regulators of methyltransferase activity. These results also link DNA methylation to sequence-specific DNA binding activities and cellular activities regulating growth control. Also, mutations in ATRX (see below) have multiple effects on methylation implicating this factor in methylation regulation.
Cis-acting regulatory factors
Information about cis-acting regulation of DNA methylation has come from studies of the imprinted loci (see below). The reproducible, parent-of-origin-specific patterns of methylation detectable at those loci provide models for identifying DNA elements that determine whether or not the locus acquires methylation. The Igf2r locus is methylated on the maternal allele which is also the expressed allele in mice. A 3 kbp intronic element termed region 2 was shown in transgenic studies to contain the imprinting center needed for imprinted methylation and expression.12 Subsequent studies identified two sequences within region 2 that regulate maternal allele-specific methylation in the post-zygotic, pre-implantation embryo. One sequence termed the DNS provides a d e n ovo methylation signal and is needed for methylation establishment of the maternal allele. A second sequence called the ADS provides an allele discrimination signal that protects the paternal allele from acquiring methylation directed by the DNS. A factor detectable in both androgenetic and gynogenetic ES cells can interact with the DNS while a factor detectable only in androgenetic ES cells interacts with the ADS.13 Presumably, the methylation patterns seen in later embryos and adults at the endogenous Igf2r locus depend upon these cis-acting elements.
The Rasgrf1 locus is methylated on the paternal allele. Expression is also paternal allele-specific in neonatal brain.14 Deletion of a repeated sequence element (40 copies of a 41-mer) 3′ of the differentially methylated domain prevented establishment of paternal allele methylation in the male germ line. Methylation patterns, once disrupted at the establishment stage, never became established in the soma.15 Interestingly, repeats and inverted repeats have been described as regulating local methylation in mouse cells16 and in plants.17
At the H19/Igf2 locus, a region 5′ to the H19 gene is methylated on the paternal allele. Deletion of 2 kbp of this region did not perturb establishment of methylation of the remaining differentially-methylated sequences, however, established methylation patterns were not efficiently maintained.18 This highlights potentially important differences between regulation of methylation establishment in the germ line and maintenance of established patterns in somatic tissue. A recent study identified Dnmt3L as a key player in the establishment of DMR methylation. Dnmt3L is expressed during gametogenesis at stages where genomic imprints are established. Mice lacking this gene do not establish methylation on the maternal allele in DMRs indicating the importance of DNMT3L for the imprinting process.19
Further clues about cis- and trans-acting factors regulating methylation in mammals may be provided by genetic screens in plants and fungi for modifiers of methylation-dependent processes.17,20,21,22 Results from these studies demonstrate conservation in plants of factors known to be involved in DNA methylation in mammals. It is likely that these systems will reveal important clues about how methylation control goes awry in human cancer.
Methylation and transcription
It is now well established that DNA methylation is involved in regulating gene transcription. This can occur by a variety of mechanisms. The interactions of several transcription factors whose binding sites contain CpG dinucleotides has been shown to be methylation-sensitive.23,24 However, methylated DNA more profoundly affects transcription by interacting with methyl-CpG-binding proteins and associated factors that alter chromatin structure.
The first two methyl-CpG-binding proteins (or protein complexes) to be identified were MeCP1 and MeCP2.25,26 MeCP2 is a single polypeptide with a methyl CpG binding domain (MBD) and a transcriptional repression domain (TRD).27 Interestingly, mutations in MeCP2 cause Rett syndrome, one of the leading causes of mental retardation and autistic behaviour.28 The MBD motif is found in four additional methyl-CpG-binding proteins (MBD1, 2, 3, and 4).29 One of these (MBD2) facilitates the binding of the multiprotein MeCP1 complex to methylated DNA.30 How proteins with methyl-CpG-binding activities repress transcription is under active study. There is evidence for a variety of complex mechanisms. One key mechanism involves MBD-mediated recruitment to methylated DNA one of two co-repressor complexes, Sin3 and Mi-2/NuRD, which in turn recruit a core histone deacetylase complex consisting of HDAC1, HDAC2 and two Rb associated, histone-binding proteins, RbAP46 and RbAP48 (31,32,33 reviewed in34,35). Additional co-repressor complexes exist, however, their role in silencing mechanisms that involve DNA methylation has not been demonstrated. HDACs remove acetyl groups from the lysine residues found at the N-termini of histone H3 and H4 (reviewed in36). Their removal results in an increase in the positive charge of the histones which is hypothesised to condense chromatin by enabling a tighter association between the histones and the negatively charged DNA. This may in turn silence transcription by limiting transcription factor binding. The Mi-2/NuRD co-repressor complex which recruits the HDAC core complex also includes factors that remodels chromatin by ATP-dependent mechanisms.37 These activities can reposition nucleosomes on DNA which may restrict interactions between the DNA and transcription factors.
This description is surely an over-simplification. First, the chromatin remodelling and histone deacetylation activities are interdependent. It has been observed that the remodelling activity of Mi-2/NuRD is required for Mi-2/NuRD-dependent deacetylation by the core HDAC complex (37 reviewed in38). Second, DNA methylation may require HDAC activity or components of the chromatin remodelling apparatus. In support of this are the observations that treatment of Neurospora crassa with the HDAC inhibitor trichostatin A (TSA) can lead to loss of DNA methylation,39 mutations in ATRX, a member of the SWI/SNF chromatin remodelling family, causes regional hyper- and hypo-methylation in the genome34 and DNMTs form complexes with HDACs.5,10,40,41 Third, adding to the varied interrelationships among silencing mechanisms is the fact that HDACs recruit sequence-specific transcriptional repressors that contribute to silencing, so transcriptional repression by HDACs may not rely solely upon DNA methylation to identify loci to be repressed. Similarly, acetylation- and methylation-mediated repression may function independently. This is supported by studies using the demethylating agent 5-aza-dC and the HDAC inhibitor, TSA. In vitro studies show that for several repressed loci, both drugs were needed for maximum de-repression – one alone would not suffice.30 Taken together, these observations highlight the complex interactions among DNA methylation, histone acetylation, chromatin remodelling and sequence-specific silencing mechanisms that collectively attenuate gene expression.
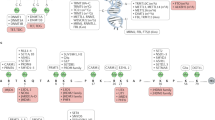
While methylation is commonly associated with transcriptional repression, there are at least two cases at imprinted loci where methylation results in transcriptional activation. At the imprinted loci H19/Igf2 and Rasgrf1, the differentially methylated domains possess enhancer-blocking activity that restricts enhancer to promoter interactions, preventing promoter activation (42,43 and Yoon et al. unpublished). Binding of the factor CTCF, which is required for the blocking activity, is methylation-sensitive (Figure 2). Methylation of the CTCF site blocks binding and restores enhancer to promoter interactions which activates expression of Igf2 or Rasgrf1. Biallelic lgf2 expression seen in cancers may involve interference with this mechanism. The mechanism by which CTCF prevents transcription may involve the binding of SIN3A and associated HDACs to an internal 11 Zn finger-containing region of CTCF and possibly additional co-repressors to the C-terminus.44
Methylation and the regulation of imprinted IGF2/H19. IGF2 and H19 are two imprinted genes located within the same chromosomal domain. IGF2 is normally transcribed from the paternal (P) allele while H19 is transcribed from the maternal (M). Imprinted expression in both genes is regulated by an imprinting center (IC) in which the maternal allele is unmethylated (open circles) and the paternal is methylated (filled circles). The imprinting center contains CTCF binding sites to which CTCF binds only if they are unmethylated. Binding of CTCF blocks the activity of the enhancer element (E) located downstream of H19 and restricts activity to H19 expression. Methylation of the CTCF binding sites prevents the binding of CTCF and allows the enhancer to activate IGF2. Biallelic expression of the CTCF binding sites in cancer results in IGF2 overexpression.
There may be additional mechanisms by which DNA methylation affects transcription. In vitro studies have shown that methylation at cytosine residues can cause several changes in DNA structure (reviewed in45). Methylation has been shown to increase the helical pitch of DNA,46 alter the rate constants for cruciform formation, lower the free energy of Z-DNA formation, and promote helix unwinding at B/Z-DNA junctions.47 If these DNA structural changes occur in vivo and affect binding of transcription-regulating factors is not known.
Regardless of the mechanisms by which methylation regulates gene expression, a large number of genes respond to methylation levels. In studies using fibroblast cells in which a Dnmt1 deletion was induced, up to 10% of over 5000 genes examined underwent significant changes in expression upon induction of the deletion.48
Compartmentalisation of the genome by DNA methylation patterns
The human genome is divided into several functionally different compartments with different base composition. Compartments include gene rich or gene poor regions, intergenic sequences, interspersed repetitive elements, and middle repetitive sequences such as the centromeric repeats. Further compartmentalisation comes through epigenetic modifications including different densities of chromatin condensation and DNA methylation patterns. CpG dinucleotides, the major target sequences for DNA methylation, are underrepresented in the genome compared to other dinucleotides. However, some GC-rich sequence stretches contain a higher frequency of CG dinucleotides. For example, CpG islands are GC-rich sequences between 0.2 and 2 kb in size and contain a higher than expected number of CpG dinucleotides (CpG to GpC ratio>0.6) and have a GC-content of more than 50% (reviewed in49). The majority of CpG islands are found in the promoter regions of genes. The majority of all CG dinucleotides in the genome are found repetitive elements such as rDNA, satellite sequences and centromeric repeats, and are usually unmethylated.
Genomic imprinting and the involvement of DNA methylation
The importance of specific DNA methylation patterns for developmentally appropriate gene expression is most clearly demonstrated for the imprinted loci. Normally, genes are expressed from both the maternal and the paternal alleles. At the imprinted loci, only the maternal or the paternal allele is expressed (for reviews see:50,51,52,53). Over 50 imprinted loci have been found in mammals (see http://cancer.otago.ac.nz/IGC/Web/home.html and http://www3.ncbi.nlm.nih.gov/Omim/ search term ‘imprint’). This restriction may be limited to specific tissues or times during development. The methylation status of the DNA surrounding an imprinted locus also displays a pattern that is unique to each allele. The locations of the differentially methylated domains or regions (DMDs or DMRs) are variable and the expressed allele may show both hypo- and/or hypermethylated domains (15,54,55 reviewed in56). These parental allele-specific methylation patterns may be established in primordial germ cells and are detectable in gametes, alternatively, they may appear in the zygote after fertilisation. Evidence indicates that the allele-specific patterns of methylation are what direct allele-specific expression.15,57 In the case of the H19/Igf2 and Rasgrf1 loci, the DMRs have enhancer blocking activity and bind CTCF in a methylation-sensitive manner42,43,58 (and Yoon unpublished result). As described earlier, CTCF bound to an unmethylated DMR represses enhancer to promoter interactions needed for Igf2 and Rasgrf1 expression and this block is relieved allowing expression when the DMRs are methylated and CTCF binding is prevented.
Other allele-specific phenomena can be observed at imprinted loci. These include differences in the timing of replication59 and chromatin structure55,60,61 of the two parental alleles. These results reveal that whatever the mechanism of genomic imprinting, it influences several DNA-based phenomena and not just gene transcription.
II. Methylation changes in cancer
For many years, genetic changes that alter primary DNA sequence were thought to be the mechanism by which critical gene activities were lost in cancer. While these largely irreversible mutations are very common in cancer, it is now well established that DNA methylation plays a significant role in loss of gene function (reviewed in62,63). Either homozygous methylation or methylation in combination with one of the genetic alterations has been described for multiple tumour suppressor genes and candidate cancer genes. Methylation can provide one of the hits postulated in Knudson's two hit hypothesis to inactivate tumour suppressor genes. Hypermethylation events in usually unmethylated CpG islands are found in a large number of cancer genes. Hypo- and hypermethylation events are found in the same tumour samples (reviewed in64) indicating a defect in the regulatory mechanisms that participate in the establishment and maintenance of methylation patterns.
Global hypomethylation in human malignancies
Methylation changes in human malignancies were first reported almost 20 years ago.65 Studies by Ehrich65 and Gama-Sosa,66 using HPLC to determine the 5′-methylcytosine content in genomic DNAs, demonstrated a reduction of the overall amount of 5′-methylcytosine relative to normal tissues. Subsequent studies demonstrated a positive correlation between the degree of hypomethylation and the increasing malignancy grade or a correlation6667,68 between tumour size and histological grade suggesting that hypomethylation may be a useful biomarker with prognostic significance.69 While global assays show that genome-wide levels of methylation decrease in many tumours, they do not provide information on where the hypomethylation occurs.
Hypermethylation in cancer
While global hypomethylation is detectable in the cancer genome, regions of hypermethylation are commonly found. Locus-specific methylation changes were initially studied using methylation sensitive restriction enzymes to digest genomic DNAs. A major breakthrough was made with the finding that sodium bisulphite treatment converts unmethylated cytosines to uracil whereas methylated cytosines remain unmodified.70 This treatment generates different sequences based on the methylation status of a gene. Subsequent PCR reactions using bisulphite-treated DNA can be performed to detect specifically the methylated sequences (MsPCR, Ms SNuPE).71,72 Alternatively, PCR products can be cloned and sequenced to identify methylated regions. A major emphasis in the past was the investigation of promoter methylation in known tumour suppressor genes. The list of genes that are found to be inactivated by DNA methylation events is growing rapidly and includes genes involved in apoptosis, angiogenesis, cell-cycle, differentiation, DNA repair, metastasis, signal transduction and transcription. Defects in a number of these genes have been identified as the underlying cause of familial cancer syndromes (reviewed in64).
The effect of DNA methylation on transcription in cancer cell lines show that hypermethylation most commonly causes gene silencing. Treatment of these cell lines with 5′–aza-2′-deoxycytidine results in demethylation of previously methylated sequences and subsequent activation of the gene under investigation.
CpG island hypermethylation
More recently, two methods were introduced that allow scanning for CpG island hypermethylation in tumour samples. Differential methylation hybridisation (DMH) is an array based approach in which CpG island sequences are spotted in high density onto nylon membranes.73 These arrays are subsequently hybridised with pools of PCR products derived from CpG islands that were amplified from genomic DNAs following BstUI digest. BstUI is a methylation sensitive restriction enzyme and its recognition sequence is frequently located in CpG island sequences. PCR products are only amplified when the target sequence is methylated and thus undigested. Array profiles from normal and tumour tissue can be compared and methylated sequences are visualised by stronger signal intensities in the tumour profile compared to the normal tissue profile. This technique was used to study methylation in breast cancer and it was shown that overall levels of CpG island methylation correlate with histological grades. Poorly differentiated tumours appeared to show more hypermethylated CpG island sequences than moderately and well differentiated tumours.74
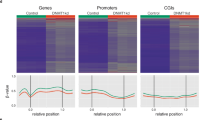
Restriction Landmark Genomic Scanning (RLGS), a two-dimensional gel-electrophoretic technique, allows the assessment of methylation patterns in up to 2000 CpG islands per gel by using rare-cutting, methylation sensitive restriction enzymes (e.g. NotI or AscI)75. RLGS profiles display unmethylated CpG islands, whereas methylated CpG islands are not displayed. Aberrant methylation in primary tumours or cancer cell lines is identified by comparing tumour RLGS profiles to profiles from matching normal DNAs. Loci aberrantly methylated in tumours are easily cloned using arrayed boundary libraries and subsequent database searches allow to link the methylation events to genes or ESTs.
Genome-wide scans for methylation changes in CpG islands showed that the frequencies of methylation are different between tumour types.75 Breast, head and neck and testicular tumours showed either no or relatively low frequencies (<1%) of methylated CpG islands. Other malignancies including acute myeloid leukemias, colon cancer and brain tumours showed an overall much higher frequency of methylation ranging up to 10% of all CpG islands tested. Patterns of methylation were not random, suggesting the existence of mechanisms that either allow the preferential methylation of certain CpG islands or a selective pressure that favours the growth of cells with specifically methylated CpG island sequences. The non-random nature of CpG island methylation patterns was underlined by the finding of several hypermethylated CpG islands in the major breakpoint cluster region for medulloblastomas on chromosome 17p11.2, suggesting a possible link between chromosomal instability in this region and hypermethylation events.76 Acute myeloid leukaemias showed a prevalence of hypermethylated CpG islands on chromosome 11, again supporting the finding that methylation patterns are not random.77 Careful inspection of the methylated target sequences allowed the identification of sequences that were methylated in several different tumour types and others that were methylated in a tumour-type specific manner. This finding is in line with reports by other groups that have found BRCA1 promoter methylation only in breast and ovarian cancers, VHL promoter methylation only in clear cell renal carcinomas and hemangioblastomas, while other tumour suppressor genes including p16, DAPK, MGMT, have been found methylated in multiple tumour types.64
Methylated genes
Both strategies, the gene-by-gene approach as well as the genome scanning approach, provided long lists of methylation targets.64,77,78,79 The significance of tumour suppressor gene inactivation in tumour progression has been already established. However, this is not the case for the genes identified in the genome scans for aberrant methylation. Genome scans may identify candidate tumour suppressor genes and genes for which no tumour suppressor function has been established. Several current examples should be highlighted. First, RLGS was used to scan hepatocellular carcinomas and identified several RLGS sequences that were methylated in the tumour samples.80 Cloning of one of these loci (spot 7) identified the promoter region of suppressor of cytokine signalling (SOCS1).81 SOCS1 is a member of the JAK/STAT pathway where by binding of SOCS1 to JAK2 phosphorylation is inhibited. Silencing of SOCS1 by promoter methylation results in constitutive activation of the JAK/STAT pathway and subsequent transactivation of target genes. The second example is bone morphogenic protein 3B (BMP3B), identified in a screen for aberrant methylation in non-small cell lung cancers.79 BMP3B is a member of the transforming growth factor-β (TGFβ) superfamily, a group of secreted polypeptides that regulate a diverse spectrum of developmental processes. Members of this superfamily signal through Ser/Thr kinase receptor which subsequently propagate signals to the SMAD pathway. Methylation of the BMP3B promoter was shown to correlate with gene silencing and could be reactivated in lung cancer cell lines by 5′-aza-2′-deoxycytidine. The effects of BMP3B silencing for tumour progression need to be demonstrated,79 however, BMP3B is located on chromosome 10q11, an area that shows loss of heterozygousity in lung cancer.
RAS effector homologue (RASSF1), a novel lung cancer tumour suppressor candidate gene, is located in a commonly deleted region of chromosome 3p. RASSF1A, the major transcript of this gene, is also silenced by DNA methylation in lung tumours.82 Subsequent reports have also demonstrated the silencing of RASSF1A by promoter methylation in several other cancer types.83,84 Re-expression of RASSF1A in lung cancer cell lines resulted in reduced numbers of colonies in colony formation assays, suppressed the anchorage independent growth and most importantly inhibited the formation of tumours in nude mice.82
Methylation profiles and clinical diagnostics
Genome scans such as RLGS allow the correlation of thousands of methylation events to clinical data. Frühwald et al. identified eight sequences in an RLGS scan that showed statistical significant correlation with survival of medulloblastoma patients.85 Surprisingly, one of these methylation events correlated with improved outcome. RLGS analysis in AML patients identified a sequence within the WIT1 gene that was methylated at a higher frequency in relapsed AML as compared to diagnostic samples thus correlating with a chemoresistant phenotype.86
Dysregulation of genomic imprinting in cancer
In addition to silencing known and candidate tumour suppressors, aberrant DNA methylation affects expression of imprinted genes. Loss of imprinting (LOI) of IGF2 and the tightly-linked H19 locus has been associated with tumorigenesis in a variety of patients. In Wilms' tumour, imprinted expression of IGF2 is commonly relaxed resulting in maternal allele expression.87,88 This is associated with increased methylation and reduced expression from the maternal copy of the tightly linked H19 locus.89,90,91 Similar patterns of LOI for either or both of these loci have been seen in patients with hepatoblastoma,92 uterine leiomyosarcomata,93 cervical carcinoma,94 renal cell carcinoma,95 rhadbomyosarcoma,96,97 gliomas98 and colorectal cancer.99,100 It is not known if LOI of IGF2 or other imprinted loci is a cause or a consequence of neoplastic transformation, however, the importance of IGF2 in human cancer is supported by the observation that patients with Beckwith-Wiedemann Syndrome (BWS) express both IGF2 alleles and are predisposed to several cancers.101,102,103
The mechanisms and methylation events underlying the LOI at H19 and IGF2 vary from tumour to tumour, however, maternal allele methylation is commonly acquired at sequences that include the H19 promoter and IGF2 enhancer blocker104,105 which can simultaneously silence H19 and activate IGF2 on the maternal chromosome42 (see Figure 2).
Imprinted loci other than H19/IGF2 have also been implicated in cancer. In neuroblastomas without N-MYC amplification, maternal-specific deletions of 1p36 have been reported.106 This region was later shown to contain the p73 tumor suppressor, a p53 homologue which is maternally-expressed.107 Silencing of the active maternal allele by methylation has been observed in acute leukaemia and Burkitt's lymphoma108 but it is possible that methylation-independent means of silencing occur in other cancers.109 p57KIP2, a cyclin dependent kinase inhibitor, is expressed primarily from the maternal allele.110 In lung cancers with 11p15 deletions, the maternal allele is predominantly deleted.111 In Wilms' tumours, it is substantially silenced,112 however, it is not clear if this results from aberrant methylation. NOEY2/ARHI is a maternally-expressed tumour suppressor commonly found to be silenced by gene deletion in breast and ovarian cancers. The methylation status of intact alleles from tumours is not known.113 In hepatocellular carcinomas, multiple imprinted loci within the 11p15 region including CDKN1C, SLC22A1L, and IGF2 genes were aberrantly silenced.114
Causes of aberrant methylation
Little is known about the mechanisms regulating DNA methylation in normal development and less is known about how these mechanisms go awry in cancer. Many correlative studies revealed increased levels of DNMTs in some cancer tissues or cells (reviewed in115), however, this was not universal.116 Furthermore, even when the correlation held, over expression of DNMTs alone was not sufficient for altered methylation patterns.117 Experimental models have supported the importance of DNMT expression levels in cancer development. In mice of strains susceptible to tumour formation, increased levels of Dnmt1 and methylated DNA were found in lungs from carcinogen-treated animals when compared to mice of resistant strains. However, in the strains used, there were many other strain-specific differences beyond Dnmt1 expression levels.118 In more easily controlled studies using Min mice which are predisposed to develop intestinal tumors, experimental reduction in Dnmt1 levels reduced tumour incidence.119 This is consistent with earlier observations that Dnmt1 has transforming activity in vitro.120
Regardless of the levels of DNMT protein, regulation of DNMT activity may be lost in cancer cells or precancerous lesions. This may be through changes in DNMT-associated factors that may regulate activity or subcellular localization. HDAC2, DMAP1 and TSG10140 and two MBDs (MBD2 and MBD3)11 have all been shown to interact with DNMT1 and a histone methyltransferase is important to DNA methylation in Neurospora.9 If any of these are important for regulating DNMT activity in mammals and whether interfering with such regulation contributes to aberrant DNA methylation in cancer is not known.
III. Perspectives
As DNA methylation is important in the regulation of gene expression one could argue that the identification of aberrant methylation patterns could serve as a biomarker with predictive values for future progression or for the response to treatment. Several groups have been interested in these approaches. In a recent study by Esteller et al. the authors describe promoter methylation in the DNA repair gene O6-methyl-guanine-DNA methyltransferase (MGMT).121 MGMT repairs alkylated sites in the DNA that otherwise would form cross-links between adjacent DNA strands. Alkylation of the DNA is the underlying principle of chemo-therapeutics such as carmustine that kill tumour cells and are used to treat patients with gliomas. Silencing of MGMT by promoter methylation was shown to correlate with the responsiveness of the gliomas to the treatment with alkylating agents. A different approach was used by Palisamo et al. in lung cancers where highly sensitive MS-PCR reactions were used to detect p16 and MGMT promoter methylation in sputum of patients up to three years prior to clinical diagnosis).122 This strategy offers the intriguing possibility of a population based screening for the detection of lung cancer. The identification of a powerful biomarker does not require the knowledge of the effect of DNA methylation on gene transcription. Several studies have identified methylated target sequences with statistically significant values.
The past years have moved the field of DNA methylation to a new level and improved our knowledge of targets affected by aberrant methylation in the cancer genome and how these changes lead to altered gene expression. Using model systems and selected genes, especially imprinted genes, it became possible to design studies that helped to explain the underlying mechanisms that regulate methylation changes. Future work will focus on several unanswered questions to determine how sequences become targets for aberrant methylation in cancer, to determine the regulatory mechanisms that malfunction in this disease and to determine the functional consequences of aberrant methylation. Identification of the factors regulating DNA methylation and demethylation may reveal novel targets for therapeutic intervention. Because DNA methylation does not alter the primary DNA sequence and is reversible, epigenetics-based therapeutics may be able to restore the activity of silenced tumor suppressors even in advanced tumours.
References
Ramsahoye BH, Biniszkiewicz D, Lyko F, Clark V, Bird AP, Jaenisch R . Non-CpG methylation is prevalent in embryonic stem cells and may be mediated by DNA methyltransferase 3a [In Process Citation] Proc Natl Acad Sci USA 2000 97: 5237–5242
Vertino PM, Yen RW, Gao J, Baylin SB . De novo methylation of CpG island sequences in human fibroblasts overexpressing DNA (cytosine-5-)-methyltransferase Mol Cell Biol 1996 16: 4555–4565
Li E, Bestor TH, Jaenisch R . Targeted mutation of the DNA methyltransferase gene results in embryonic lethality Cell 1992 69: 915–926
Okano M, Bell DW, Haber DA, Li E . DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development Cell 1999 99: 247–257
Fuks F, Burgers WA, Godin N, Kasai M, Kouzarides T . Dnmt3a binds deacetylases and is recruited by a sequence-specific repressor to silence transcription EMBO J 2001 20: 2536–2544
Robertson KD, Uzvolgyi E, Liang G et al. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: coordinate mRNA expression in normal tissues and overexpression in tumors Nucleic Acids Res 1999 27: 2291–2298
Howell CY, Bestor TH, Ding F et al. Genomic imprinting disrupted by a maternal effect mutation in the Dnmt1 gene Cell 2001 104: 829–838
Oswald J, Engemann S, Lane N et al. Active demethylation of the paternal genome in the mouse zygote [In Process Citation] Curr Biol 2000 10: 475–478
Tamaru H, Selker EU . A histone H3 methyltransferase controls DNA methylation in Neurospora crassa Nature 2001 414: 277–283
Robertson KD, Ait-Si-Ali S, Yokochi T, Wade PA, Jones PL, Wolffe AP . DNMT1 forms a complex with Rb, E2F1 and HDAC1 and represses transcription from E2F-responsive promoters Nat Genet 2000 25: 338–342
Tatematsu KI, Yamazaki T, Ishikawa F . MBD2-MBD3 complex binds to hemi-methylated DNA and forms a complex containing DNMT1 at the replication foci in late S phase Genes Cells 2000 5: 677–688
Wutz A, Smrzka OW, Schweifer N, Schellander K, Wagner EF, Barlow DP . Imprinted expression of the Igf2r gene depends on an intronic CpG island [see comments] Nature 1997 389: 745–749
Birger Y, Shemer R, Perk J, Razin A . The imprinting box of the mouse Igf2r gene Nature 1999 397: 84–88
Plass C, Shibata H, Kalcheva I et al. Identification of Grf1 on mouse chromosome 9 as an imprinted gene by RLGS-M Nat Genet 1996 14: 106–109
Yoon B-J, Herman H, Sikora A . Regulation of DNA methylation of Rasgrf1 Nature Genet 2002 30: 92–96
Yates PA, Burman RW, Mummaneni P, Krussel S, Turker MS . Tandem B1 elements located in a mouse methylation center provide a target for de novo DNA methylation J Biol Chem 1999 274: 36357–36361
Luff B, Pawlowski L, Bender J . An inverted repeat triggers cytosine methylation of identical sequences in Arabidopsis Mol Cell 1999 3: 505–511
Thorvaldsen JL, Duran KL, Bartolomei MS . Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2 Genes Dev 1998 12: 3693–3702
Bourc'his D, Xu GL, Lin CS, Bollman B, Bestor TH . Dnmt3L and the Establishment of Maternal Genomic Imprints Science 2001 22: 22
Pelissier T, Wassenegger M . A DNA target of 30 bp is sufficient for RNA-directed DNA methylation Rna 2000 6: 55–65
Cogoni C, Irelan JT, Schumacher M, Schmidhauser TJ, Selker EU, Macino G . Transgene silencing of the al-1 gene in vegetative cells of Neurospora is mediated by a cytoplasmic effector and does not depend on DNA-DNA interactions or DNA methylation EMBO J 1996 15: 3153–3163
Selker EU . Gene silencing: repeats that count Cell 1999 97: 157–160
Watt F, Molloy PL . Cytosine methylation prevents binding to DNA of a HeLa cell transcription factor required for optimal expression of the adenovirus major late promoter Genes Dev 1988 2: 1136–1143
Kovesdi I, Reichel R, Nevins JR . Role of an adenovirus E2 promoter binding factor in E1A-mediated coordinate gene control Proc Natl Acad Sci USA 1987 84: 2180–2184
Lewis JD, Meehan RR, Henzel WJ et al. Purification, sequence, and cellular localization of a novel chromosomal protein that binds to methylated DNA Cell 1992 69: 905–914
Meehan RR, Lewis JD, McKay S, Kleiner EL, Bird AP . Identification of a mammalian protein that binds specifically to DNA containing methylated CpGs Cell 1989 58: 499–507
Nan X, Ng HH, Johnson CA et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex [see comments] Nature 1998 393: 386–389
Amir RE, Van den Veyver IB, Wan M, Tran CQ, Francke U, Zoghbi HY . Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2 Nat Genet 1999 23: 185–188
Hendrich B, Bird A . Identification and characterization of a family of mammalian methyl-CpG binding proteins Mol Cell Biol 1998 18: 6538–6547
Ng HH, Zhang Y, Hendrich B et al. MBD2 is a transcriptional repressor belonging to the MeCP1 histone deacetylase complex Nat Genet 1999 23: 58–61
Laherty CD, Yang WM, Sun JM, Davie JR, Seto E, Eisenman RN . Histone deacetylases associated with the mSin3 corepressor mediate mad transcriptional repression Cell 1997 89: 349–356
Nagy L, Kao HY, Chakravarti D et al. Nuclear receptor repression mediated by a complex containing SMRT, mSin3A, and histone deacetylase Cell 1997 89: 373–380
Zhang Y, Ng HH, Erdjument-Bromage H, Tempst P, Bird A, Reinberg D . Analysis of the NuRD subunits reveals a histone deacetylase core complex and a connection with DNA methylation Genes Dev 1999 13: 1924–1935
Gibbons RJ, McDowell TL, Raman S et al. Mutations in ATRX, encoding a SWI/SNF-like protein, cause diverse changes in the pattern of DNA methylation Nat Genet 2000 24: 368–371
Knoepfler PS, Eisenman RN . Sin meets NuRD and other tails of repression Cell 1999 99: 447–450
Kuo MH, Allis CD . Roles of histone acetyltransferases and deacetylases in gene regulation Bioessays 1998 20: 615–626
Guschin D, Wade PA, Kikyo N, Wolffe AP . ATP-Dependent histone octamer mobilization and histone deacetylation mediated by the Mi-2 chromatin remodeling complex Biochemistry 2000 39: 5238–5245
Tyler JK, Kadonaga JT . The ‘dark side’ of chromatin remodeling: repressive effects on transcription Cell 1999 99: 443–446
Selker EU . Trichostatin A causes selective loss of DNA methylation in Neurospora Proc Natl Acad Sci USA 1998 95: 9430–9435
Rountree MR, Bachman KE, Baylin SB . DNMT1 binds HDAC2 and a new co-repressor, DMAP1, to form a complex at replication foci Nat Genet 2000 25: 269–277
Fuks F, Burgers WA, Brehm A, Hughes-Davies L, Kouzarides T . DNA methyltransferase Dnmt1 associates with histone deacetylase activity Nat Genet 2000 24: 88–91
Hark AT, Schoenherr CJ, Katz DJ, Ingram RS, Levorse JM, Tilghman SM . CTCF mediates methylation-sensitive enhancer-blocking activity at the H19/Igf2 locus [see comments] Nature 2000 405: 486–489
Bell AC, Felsenfeld G . Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene [see comments] Nature 2000 405: 482–485
Lutz M, Burke LJ, Barreto G et al. Transcriptional repression by the insulator protein CTCF involves histone deacetylases Nucleic Acids Res 2000 28: 1707–1713
Zacharias W . Methylation of cytosine influences the DNA structure Exs 1993 64: 27–38
Gruenbaum Y, Cedar H, Razin A . Substrate and sequence specificity of a eukaryotic DNA methylase Nature 1982 295: 620–622
Zacharias W, Caserta M, O'Connor TR, Larson JE, Wells RD . Cytosine methylation as an effector of right-handed to left-handed DNA structural transitions Gene 1988 74: 221–224
Jackson-Grusby L, Beard C, Possemato R et al. Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation [In Process Citation] Nat Genet 2001 27: 31–39
Antequera F, Bird A . Number of CpG islands and genes in human and mouse Proc Natl Acad Sci USA 1993 90: 11995–11999
Bartolomei MS, Tilghman SM . Genomic imprinting in mammals Annu Rev Genet 1997 31: 493–525
Feil R, Khosla S . Genomic imprinting in mammals: an interplay between chromatin and DNA methylation? Trends Genet 1999 15: 431–435
Pfeifer K . Mechanisms of genomic imprinting Am J Hum Genet 2000 67: 777–787
Reik W, Walter J . Genomic imprinting: parental influence on the genome Nat Rev Genet 2001 2: 21–32
Stoger R, Kubicka P, Liu CG et al. Maternal-specific methylation of the imprinted mouse Igf2r locus identifies the expressed locus as carrying the imprinting signal Cell 1993 73: 61–71
Bartolomei MS, Webber AL, Brunkow ME, Tilghman SM . Epigenetic mechanisms underlying the imprinting of the mouse H19 gene Genes Dev 1993 7: 1663–1673
Mann JR, Szabo PE, Reed MR, Singer-Sam J . Methylated DNA sequences in genomic imprinting Crit Rev Eukaryot Gene Expr 2000 10: 241–257
Li E, Beard C, Jaenisch R . Role for DNA methylation in genomic imprinting [see comments] Nature 1993 366: 362–365
Kanduri C, Pant V, Loukinov D et al. Functional association of CTCF with the insulator upstream of the H19 gene is parent of origin-specific and methylation-sensitive Curr Biol 2000 10: 853–856
Simon I, Tenzen T, Reubinoff BE, Hillman D, McCarrey JR, Cedar H . Asynchronous replication of imprinted genes is established in the gametes and maintained during development Nature 1999 401: 929–932
Feil R, Boyano MD, Allen ND, Kelsey G . Parental chromosome-specific chromatin conformation in the imprinted U2af1-rs1 gene in the mouse J Biol Chem 1997 272: 20893–20900
Shibata H, Yoshino K, Sunahara S et al. Inactive allele-specific methylation and chromatin structure of the imprinted gene U2af1-rs1 on mouse chromosome 11 Genomics 1996 35: 248–252
Jones PA, Laird PW . Cancer epigenetics comes of age Nat Genet 1999 21: 163–167
Baylin SB, Herman JG . DNA hypermethylation in tumorigenesis: epigenetics joins genetics [In Process Citation] Trends Genet 2000 16: 168–174
Costello JF, Plass C . Methylation matters J Med Genet 2001 38: 285–303
Ehrlich M, Gama-Sosa MA, Huang LH et al. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells Nucleic Acids Res 1982 10: 2709–2721
Gama-Sosa MA, Wang RY, Kuo KC, Gehrke CW, Ehrlich M . The 5-methylcytosine content of highly repeated sequences in human DNA Nucleic Acids Res 1983 11: 3087–3095
Cravo M, Pinto R, Fidalgo P et al. Global DNA hypomethylation occurs in the early stages of intestinal type gastric carcinoma Gut 1996 39: 434–438
Kim YI, Giuliano A, Hatch KD et al. Global DNA hypomethylation increases progressively in cervical dysplasia and carcinoma Cancer 1994 74: 893–899
Soares J, Pinto AE, Cunha CV et al. Global DNA hypomethylation in breast carcinoma: correlation with prognostic factors and tumor progression Cancer 1999 85: 112–118
Clark SJ, Harrison J, Paul CL, Frommer M . High sensitivity mapping of methylated cytosines Nucleic Acids Res 1994 22: 2990–2997
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB . Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands Proc Natl Acad Sci U S A 1996 93: 9821–9826
Gonzalgo ML, Jones PA . Rapid quantitation of methylation differences at specific sites using methylation-sensitive single nucleotide primer extension (Ms-SNuPE) Nucleic Acids Res 1997 25: 2529–2531
Huang TH, Perry MR, Laux DE . Methylation profiling of CpG islands in human breast cancer cells Hum Mol Genet 1999 8: 459–470
Yan PS, Perry MR, Laux DE, Asare AL, Caldwell CW, Huang TH . CpG island arrays: an application toward deciphering epigenetic signatures of breast cancer Clin Cancer Res 2000 6: 1432–1438
Costello JF, Fruhwald MC, Smiraglia DJ et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns [see comments] Nat Genet 2000 24: 132–138
Fruhwald MC, O'Dorisio MS, Dai Z et al. Aberrant hypermethylation of the major breakpoint cluster region in 17p11.2 in medulloblastomas but not supratentorial PNETs Genes Chromosomes Cancer 2001 30: 38–47
Rush LJ, Dai Z, Smiraglia DJ, Gao X et al. Novel methylation targets in de novo acute myeloid leukemia with prevalence of chromosome 11 loci Blood 2001 97: 3226–3233
Smiraglia DJ, Rush LJ, Fruhwald MC et al. Excessive CpG island hypermethylation in cancer cell lines versus primary human malignancies Hum Mol Genet 2001 10: 1413–1419
Dai Z, Lakshmanan RR, Zhu W-G et al. Global methylation profiling of lung cancer identifies novel methylated genes Neoplasia 2001 3: 314–323
Nagai H, Ponglikitmongkol M, Mita E et al. Aberration of genomic DNA in association with human hepatocellular carcinomas detected by 2-dimensional gel analysis Cancer Res 1994 54: 1545–1550
Yoshikawa H, Matsubara K, Qian GS et al. SOCS-1, a negative regulator of the JAK/STAT pathway, is silenced by methylation in human hepatocellular carcinoma and shows growth-suppression activity Nat Genet 2001 28: 29–35
Dammann R, Li C, Yoon JH, Chin PL, Bates S, Pfeifer GP . Epigenetic inactivation of a RAS association domain family protein from the lung tumour suppressor locus 3p21.3 Nat Genet 2000 25: 315–319
Dreijerink K, Braga E, Kuzmin I et al. The candidate tumor suppressor gene, RASSF1A, from human chromosome 3p21.3 is involved in kidney tumorigenesis Proc Natl Acad Sci USA 2001 98: 7504–7509
Agathanggelou A, Honorio S, Macartney DP et al. Methylation associated inactivation of RASSF1A from region 3p21.3 in lung, breast and ovarian tumours Oncogene 2001 20: 1509–1518
Fruhwald MC, O'Dorisio MS, Dai Z et al. Aberrant promoter methylation of previously unidentified target genes is a common abnormality in medulloblastomas–implications for tumor biology and potential clinical utility Oncogene 2001 20: 5033–5042
Plass C, Yu F, Yu L et al. Restriction landmark genome scanning for aberrant methylation in primary refractory and relapsed acute myeloid leukemia; involvement of the WIT-1 gene Oncogene 1999 18: 3159–3165
Rainier S, Johnson LA, Dobry CJ, Ping AJ, Grundy PE, Feinberg AP . Relaxation of imprinted genes in human cancer Nature 1993 362: 747–749
Ogawa O, Eccles MR, Szeto J et al. Relaxation of insulin-like growth factor II gene imprinting implicated in Wilms' tumour Nature 1993 362: 749–751
Moulton T, Crenshaw T, Hao Y et al. Epigenetic lesions at the H19 locus in Wilms' tumour patients Nat Genet 1994 7: 440–447
Steenman MJ, Rainier S, Dobry CJ, Grundy P, Horon IL, Feinberg AP . Loss of imprinting of IGF2 is linked to reduced expression and abnormal methylation of H19 in Wilms' tumour [published erratum appears in Nat Genet 1994 Oct;8(2):203] Nat Genet 1994 7: 433–439
Taniguchi T, Sullivan MJ, Ogawa O, Reeve AE . Epigenetic changes encompassing the IGF2/H19 locus associated with relaxation of IGF2 imprinting and silencing of H19 in Wilms tumor Proc Natl Acad Sci USA 1995 92: 2159–2163
Rainier S, Dobry CJ, Feinberg AP . Loss of imprinting in hepatoblastoma Cancer Res 1995 55: 1836–1838
Vu TH, Yballe C, Boonyanit S, Hoffman AR . Insulin-like growth factor II in uterine smooth-muscle tumors: maintenance of genomic imprinting in leiomyomata and loss of imprinting in leiomyosarcomata J Clin Endocrinol Metab 1995 80: 1670–1676
Douc-Rasy S, Barrois M, Fogel S et al. High incidence of loss of heterozygosity and abnormal imprinting of H19 and IGF2 genes in invasive cervical carcinomas. Uncoupling of H19 and IGF2 expression and biallelic hypomethylation of H19 Oncogene 1996 12: 423–430
Oda H, Kume H, Shimizu Y, Inoue T, Ishikawa T . Loss of imprinting of igf2 in renal-cell carcinomas Int J Cancer 1998 75: 343–346
Pedone PV, Tirabosco R, Cavazzana AO et al. Mono- and bi-allelic expression of insulin-like growth factor II gene in human muscle tumors Hum Mol Genet 1994 3: 1117–1121
Zhan S, Shapiro DN, Helman LJ . Activation of an imprinted allele of the insulin-like growth factor II gene implicated in rhabdomyosarcoma J Clin Invest 1994 94: 445–448
Uyeno S, Aoki Y, Nata M et al. IGF2 but not H19 shows loss of imprinting in human glioma Cancer Res 1996 56: 5356–5359
Cui H, Horon IL, Ohlsson R, Hamilton SR, Feinberg AP . Loss of imprinting in normal tissue of colorectal cancer patients with microsatellite instability Nat Med 1998 4: 1276–1280
Takano Y, Shiota G, Kawasaki H . Analysis of genomic imprinting of insulin-like growth factor 2 in colorectal cancer Oncology 2000 59: 210–216
Weksberg R, Shen DR, Fei YL, Song QL, Squire J . Disruption of insulin-like growth factor 2 imprinting in Beckwith-Wiedemann syndrome Nat Genet 1993 5: 143–150
Ohlsson R, Nystrom A, Pfeifer-Ohlsson S et al. IGF2 is parentally imprinted during human embryogenesis and in the Beckwith-Wiedemann syndrome Nat Genet 1993 4: 94–97
Mannens M, Hoovers JM, Redeker E et al. Parental imprinting of human chromosome region 11p15.3-pter involved in the Beckwith-Wiedemann syndrome and various human neoplasia Eur J Hum Genet 1994 2: 3–23
Nakagawa H, Chadwick RB, Peltomaki P, Plass C, Nakamura Y, de La Chapelle A . Loss of imprinting of the insulin-like growth factor II gene occurs by biallelic methylation in a core region of H19-associated CTCF-binding sites in colorectal cancer Proc Natl Acad Sci USA 2001 98: 591–596
Takai D, Gonzales FA, Tsai YC, Thayer MJ, Jones PA . Large scale mapping of methylcytosines in CTCF-binding sites in the human H19 promoter and aberrant hypomethylation in human bladder cancer Hum Mol Genet 2001 10: 2619–2626
Caron H, Peter M, van Sluis P et al. Evidence for two tumour suppressor loci on chromosomal bands 1p35-36 involved in neuroblastoma: one probably imprinted, another associated with N-myc amplification Hum Mol Genet 1995 4: 535–539
Kaghad M, Bonnet H, Yang A et al. Monoallelically expressed gene related to p53 at 1p36, a region frequently deleted in neuroblastoma and other human cancers Cell 1997 90: 809–819
Corn PG, Kuerbitz SJ, van Noesel MM et al. Transcriptional silencing of the p73 gene in acute lymphoblastic leukemia and Burkitt's lymphoma is associated with 5′ CpG island methylation Cancer Res 1999 59: 3352–3356
Banelli B, Casciano I, Romani M . Methylation-independent silencing of the p73 gene in neuroblastoma Oncogene 2000 19: 4553–4556
Matsuoka S, Thompson JS, Edwards MC et al. Imprinting of the gene encoding a human cyclin-dependent kinase inhibitor, p57KIP2, on chromosome 11p15 Proc Natl Acad Sci USA 1996 93: 3026–3030
Kondo M, Matsuoka S, Uchida K et al. Selective maternal-allele loss in human lung cancers of the maternally expressed p57KIP2 gene at 11p15.5 Oncogene 1996 12: 1365–1368
Thompson JS, Reese KJ, DeBaun MR, Perlman EJ, Feinberg AP . Reduced expression of the cyclin-dependent kinase inhibitor gene p57KIP2 in Wilms' tumor Cancer Res 1996 56: 5723–5727
Yu Y, Xu F, Peng H et al. NOEY2 (ARHI), an imprinted putative tumor suppressor gene in ovarian and breast carcinomas Proc Natl Acad Sci USA 1999 96: 214–219
Schwienbacher C, Gramantieri L, Scelfo R et al. Gain of imprinting at chromosome 11p15: A pathogenetic mechanism identified in human hepatocarcinomas Proc Natl Acad Sci USA 2000 97: 5445–5449
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP . Alterations in DNA methylation: a fundamental aspect of neoplasia Adv Cancer Res 1998 72: 141–196
Eads CA, Danenberg KD, Kawakami K, Saltz LB, Danenberg PV, Laird PW . CpG island hypermethylation in human colorectal tumors is not associated with DNA methyltransferase overexpression Cancer Res 1999 59: 2302–2306
Nass SJ, Ferguson AT, El-Ashry D, Nelson WG, Davidson NE . Expression of DNA methyl-transferase (DMT) and the cell cycle in human breast cancer cells Oncogene 1999 18: 7453–7461
Belinsky SA, Nikula KJ, Baylin SB, Issa JP . Increased cytosine DNA-methyltransferase activity is target-cell-specific and an early event in lung cancer Proc Natl Acad Sci USA 1996 93: 4045–4050
Laird PW, Jackson-Grusby L, Fazeli A et al. Suppression of intestinal neoplasia by DNA hypomethylation Cell 1995 81: 197–205
Wu J, Issa JP, Herman J, Bassett DE Jr, Nelkin BD, Baylin SB . Expression of an exogenous eukaryotic DNA methyltransferase gene induces transformation of NIH 3T3 cells Proc Natl Acad Sci USA 1993 90: 8891–8895
Esteller M, Garcia-Foncillas J, Andion E et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents N Engl J Med 2000 343: 1350–1354
Palmisano WA, Divine KK, Saccomanno G et al. Predicting lung cancer by detecting aberrant promoter methylation in sputum Cancer Res 2000 60: 5954–5958
Acknowledgements
We thank LJ Rush, DJ Smiraglia, WA Held for helpful comments and honest criticism. We apologise to those authors whose work could not be cited due to restrictions in length of the review. This work was supported in part by grants P30 CA16058, R21 CA80912, RO1 GM58269, R01 DE13123 (C Plass), and P30 CA 16056 and R01 EY11279 (PD Soloway).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Plass, C., Soloway, P. DNA methylation, imprinting and cancer. Eur J Hum Genet 10, 6–16 (2002). https://doi.org/10.1038/sj.ejhg.5200768
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200768
Keywords
This article is cited by
-
Increased copy number of imprinted genes in the chromosomal region 20q11-q13.32 is associated with resistance to antitumor agents in cancer cell lines
Clinical Epigenetics (2022)
-
Association of expression of epigenetic molecular factors with DNA methylation and sensitivity to chemotherapeutic agents in cancer cell lines
Clinical Epigenetics (2021)
-
Identifying regulators of parental imprinting by CRISPR/Cas9 screening in haploid human embryonic stem cells
Nature Communications (2021)
-
DNA Methylation of T1R1 Gene in the Vegetarian Adaptation of Grass Carp Ctenopharyngodon idella
Scientific Reports (2018)
-
Genome-wide DNA methylation changes associated with olfactory learning and memory in Apis mellifera
Scientific Reports (2017)