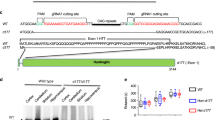
Abstract
At least five adult–onset neurodegenerative diseases, including Huntington disease (HD), and dentatorubral–pallidoluysian atrophy (DRPLA) are produced by genes containing a variably increased CAC repeat within the coding region1–4. The size range of the repeats is similar in all diseases; unaffected individuals have fewer than 30 CAG repeats, whereas affected patients usually have more than 40 repeats. The size of the inherited CAG repeat correlates with the severity and age of disease onset1,5–7. The CAG triplet repeat produces a polyglutamine domain in the expressed proteins3,8–10. All of these diseases are inherited in a dominant fashion, and a pathologic gain of function in gene carriers has been proposed. We sought to identify proteins in the brain that selectively interact with polyglutamine–domain proteins, hypothesizing that the polyglutamine domain may determine protein–protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Koide, R. et al. Unstable expansion of CAG repeat in hereditary dentatorubralpallidoluysian atrophy (DRPLA). Nature Genet. 6, 9–13 (1994).
Nagafuchi, S. et al. Dentatorubral and pallidoluysian atrophy expansion of an unstable CAG trinucleotide on chromosome 12p. Nature Genet. 6, 14–18 (1994).
Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Willems, P.J. Dynamic mutations hit double figures. Nature Genet. 8, 213–215 (1994).
Andrew, S.E. et al. The relationship between trinucleotide (CAG) repeat length and clinical features of Huntington's disease. Nature Genet. 4, 398–403 (1993).
Ranum, L.P.W. et al. Molecular and clinical correlations in spinocerebellar ataxia type I—evidence for familial effects on the age at onset. Am. J. Hum. Genet. 55, 244–252 (1994).
Kawaguchi, Y. et al. CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nature Genet. 8, 221–227 (1994).
Nagafuchi, S. et al. Structure and expression of the gene responsible for the triplet repeat disorder, dentatorubral and pallidoluysian atrophy (DRPLA). Nature Genet. 8, 177–182 (1994).
Jou, Y.S. & Myers, R.M. Evidence from antibody studies that the CAG repeat in the Huntington disease gene is expressed in the protein. Hum. Mol. Genet. 4, 465–469 (1995).
La Spada, A.R., Wilson, E.M., Lubahn, D.B., Harding, A.E. & Fischbeck, K.H. Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 352, 77–79 (1991).
Yazawa, I. et al. Abnormal gene product identified in hereditary dentatorubralpallidoluysian atrophy (DRPLA) brain. Nature Genet. 10, 99–103 (1995).
Meyer-Siegler, K. et al. A human nuclear uracil DNA glycosylase is the 37-kDa subunit of glyceraldehyde-3-phosphate dehydrogenase. Proc. Natl. Acad. Sci. USA 88, 8460–8464 (1991).
Beal, M.F. Does impairment of energy metabolism result in excitotoxic neuronal death in neurodegenerative illnesses? Ann. Neurol. 31, 119–130 (1992).
Nagy, E. & Rigby, W.F. Glyceraldehyde-3-phosphate dehydrogenase selectively binds AU-rich RNA in the NAD(+)-binding region (Rossmann fold). J. Biol. Chem. 270, 2755–2763 (1995).
Nakai, A., Satoh, M., Hirayoshi, K. & Nagata, K. Identification of the ATP-binding heat-inducible protein of MR = 37,000 as glyceraldehyde-3-phosphate dehydrogenase. Biochem. Biophys. Res. Commun. 176, 59–64 (1991).
Zeng, F.Y., Gerke, V. & Gabius, H.J. Identification of annexin II, annexin VI and glyceraldehyde-3-phosphate dehydrogenase as calcyclin-binding proteins in bovine heart. Int. J. Blochem. 25, 1019–1027 (1993).
Mejean, C., Pons, F., Benyamin, Y. & Roustan, C. Antigenic probes locate binding sites for the glycolytic enzymes glyceraldehyde-3-phosphate dehydrogenase, aldolase and phosphofructokinase on the actin monomer in microfilaments. Biochem. J. 264, 671–677 (1989).
Somers, M., Engelborghs, Y. & Baert, J. Analysis of the binding of glyceralde-hyde-3-phosphate dehydrogenase to microtubules, the mechanism of bundle formation and the linkage effect. Eur. J. Biochem. 193, 437–444 (1990).
Schulze, H. et al. Rat brain glyceraldehyde-3-phosphate dehydrogenase interacts with the recombinant cytoplasmic domain of Alzheimer's beta-amyloid precursor protein. J. Neurochem. 60, 1915–1922 (1993).
Perutz, M.F., Johnson, T., Suzuki, M. & Finch, J.T. Glutamine repeats as polar zippers: Their possible role in inherited neurodegenerative diseases. Proc. Natl. Acad. Sci. USA 91, 5355–5358 (1994).
Stott, K., Blackburn, J.M., Butler, P.J.G. & Perutz, M. Incorporation of glutamine repeats makes protein oligomerize: Implications for neurodegenerative diseases. Proc. Natl. Acad. Sci. USA 92 6509–6513 (1995).
Roses, A.D. Alzheimer's disease as a model of molecular gerontology. J. Natl. Inst. Health Res. 7 51–57 (1995).
Trottier, Y. et al. Cellular localization of the Huntington's disease protein and discrimination of the normal and mutated form. Nature Genet. 10, 104–110 (1995).
Sharp, A.H. et al. Widespread expression of Huntingtons disease gene (IT15) protein product. Neuron 14, 1065–1074 (1995).
DiFiglia, M. et al. Huntingtin is a cytoplasmic protein associated with vesicles in human and rat brain neurons. Neuron 14, 1075–1081 (1995).
van Tuinen, E. & Riezman, H. Immunolocalization of glyceraldehyde-3-phosphate dehydrogenase, hexokinase, and carboxypeptidase Y in yeast cells at the ultrastructural level. J. Histochem. Cytochem. 35, 327–333 (1987).
Minaschek, G., Groschel-Stewart, U., Blum, S. & Bereiter-Hahn, J. Microcompartmentation of glycolytic enzymes in cultured cells. Eur. J. Cell Biol. 58, 418–428 (1992).
Ronai, Z. Glycolytic enzymes as DNA binding proteins. Int. J. Biochem. 25, 1073–1076 (1993).
Morgenegg, G. et al. Glyceraldehyde-3-phosphate dehydrogenase is a nonhistone protein and a possible activator of transcription in neurons. J. Neurochem. 47 54–62 (1986).
Knull, H.R. & Fillmore, S.J. Glycolytic enzyme levels in synaptosomes. Comp. Biochem. Physiol. [B] 81, 349–351 (1985).
Hoogeveen, A.T. et al. Characterization and localization of the Huntington disease gene product. Hum. Mol. Genet. 2 2069–2073 (1993).
Servadio, A. et al. Expression analysis of the ataxin-1 protein in tissues from normal and spinocerebellar ataxia type 1 individuals. Nature Genet. 10, 94–98 (1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Burke, J., Enghild, J., Martin, M. et al. Huntingtin and DRPLA proteins selectively interact with the enzyme GAPDH. Nat Med 2, 347–350 (1996). https://doi.org/10.1038/nm0396-347
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nm0396-347
This article is cited by
-
Amelioration for an ignored pitfall in reference gene selection by considering the mean expression and standard deviation of target genes
Scientific Reports (2022)
-
Glyceraldehyde-3-Phosphate Dehydrogenase Facilitates Macroautophagic Degradation of Mutant Huntingtin Protein Aggregates
Molecular Neurobiology (2021)
-
Global Proteomic Profile Integrated to Quantitative and Morphometric Assessment of Enteric Neurons: Investigation of the Mechanisms Involved in the Toxicity Induced by Acute Fluoride Exposure in the Duodenum
Neurotoxicity Research (2021)
-
Evaluation of reference genes for gene expression studies in mouse and N2a cell ischemic stroke models using quantitative real-time PCR
BMC Neuroscience (2018)
-
Reduced cell size, chromosomal aberration and altered proliferation rates are characteristics and confounding factors in the STHdh cell model of Huntington disease
Scientific Reports (2017)