Article Text
Abstract
Background: X chromosome rearrangements defined a critical region for premature ovarian failure (POF) that extended for >15 Mb in Xq. It has been shown previously that the region could be divided into two functionally distinct portions and suggested that balanced translocations interrupting its proximal part, critical region 1 (CR1), could be responsible for POF through downregulation of ovary expressed autosomal genes translocated to the X chromosome.
Results and conclusion: This study reports that such position effect can indeed be demonstrated by analysis of breakpoint regions in somatic cells of POF patients and by the finding that CR1 has a highly heterochromatic organisation, very different from that of the euchromatic autosomal regions involved in the rearrangements. The chromatin organisation of the POF CR1 is likely to be responsible for the epigenetic modifications observed in POF patients. The characteristics of CR1 and its downregulation in oocytes may very well explain its role in POF and the frequency of the POF phenotype in chromosomal rearrangements involving Xq. This study also demonstrates a large and evolutionary conserved domain of the long arm of the X chromosome, largely corresponding to CR1, that may have structural or functional roles, in oocyte maturation or in X chromosome inactivation.
Statistics from Altmetric.com
Many rearrangements and monosomies of the X chromosome are associated with female infertility. Primary amenorrhoea and streak ovaries are consistent characteristics of X monosomy, in Turner syndrome,1 2 while premature ovarian failure (POF, OMIM 311360 and 300511), a disorder characterised by amenorrhoea and elevated serum gonadotropin values before the age of 40 years, is associated with partial monosomies and X;autosome balanced translocations.3 4 5 Cytogenetic and molecular analysis of chromosomal rearrangements has defined a critical region for POF on the long arm of the X chromosome from Xq13.3 to q26/27.6 7 Ovary expressed genes, that are significantly enriched on the mammalian X,8 could be involved in the phenotype.
We have shown that the aetiology of X linked POF may be more complex,7 9 as the POF critical region could be split into two functionally different portions, critical region 1 and 2 (CR1 and CR2). Most breakpoints in X;autosome balanced translocations in POF patients were clustered in CR1. However, only deletions involving CR2 were found to be associated with POF, while large interstitial deletions of CR1 were not. These findings suggested that POF CR1 may not carry genes for ovarian function as previously hypothesised and that such genes should be present only in CR2. Indeed, mapping of several breakpoints in the region failed to identify ovary expressed genes that could be responsible for POF. Such genes were found at the autosomal breakpoints, in six cases studied.9
Based on our data we suggested that POF CR1 may have an ovary specific conformation that will result in a position effect on autosomal genes when involved in balanced translocations. Our suggestion was strengthened by the observation that genes in the POF critical region undergo a global downregulation of expression in oocytes, compared to autosomal genes, that is not found elsewhere along the X chromosome.9 10 The downregulation of X linked gene expression in oocytes is unique as all other cell types and tissues analysed appeared instead to be upregulated in respect to the autosomes,11 possibly to reach an equal level of gene expression between the X chromosome and the autosomes.12 In oocytes, where two active X chromosomes are present, the same mechanisms may not be required, and other molecular changes, possibly essential to go through the long meiotic prophase, may become more relevant. Several mouse mutants carrying knock out (KO) in genes for chromatin modifications13 14 presented well defined meiotic defects in oocytes and/or spermatids that demonstrated the importance of a proper chromatin modifications, chromosomal structure and control of gene expression during meiosis. Based on the available evidence we proposed that oocyte expressed genes translocated to the critical region of the active X chromosome may undergo downregulation of their expression driven by the flanking X chromosome, similar to the mechanisms described for Drosophila position effect variegation.15 16 This may result in the unique and very peculiar association of the POF phenotype to X;autosome balanced translocations in CR1.
We present here a characterisation of CR1 in somatic tissues and in the ovary, in human and mouse, that indicates that CR1 is a highly heterochromatic region of the genome and indeed very different from the autosomes involved in POF associated balanced translocations; it may thus alter the epigenetic modifications of autosomal genes through a position effect detectable to some extent also in somatic cells.
Materials and methods
Cell lines and fluorescence in situ hybridisation mapping
Patients, lymphoblastoid cell lines and fluorescence in situ hybridisation (FISH) mapping have been previously described.7
Chromatin preparation and immunoprecipitation
Lymphoblastoid cell lines were fixed in 1% formaldehyde, as described by the Jenuwein laboratory (http://www.epigenome-noe.net/researchtools/protocol.php?protid = 9#ref_nav). The reaction was quenched with 125 mM glycine and the cells resuspended in lysis buffer (1% SDS, 10 mM EDTA pH8, 50 mM Tris-HCl pH 8.1, 1 mM PMSF (phenylmethylsulphonyl fluoride) and sonicated with a XL2020 sonicator microtip (Misonix Inc, Farmingdale, New York, USA) at 20% power for 150 s, to break DNA in chromatin to a size between 500–2000 bp. Chromatin was diluted 10 times in DB buffer (1% triton, 2 mM EDTA pH 8, 150 mM NaCl, 20 mM Tris-HCl pH 8.1, 1 mM PMSF). 100/200 μg of chromatin was immunoprecipitated with 2 μl of antibody specific for AcH3 (Upstate Biotechnology, Billerica, Massachusetts, USA) (UB 06–559), 5 μl AcH4 (UB 06–866) and 5 μl 2MK4H3 (UB 07–030), overnight at 4°C and then mixed with 20 μl of protein A sepharose beads (Amersham, UK) saturated overnight at 4°C with 500 μg of salmon sperm DNA and 100 μg of BSA, and incubated for 3 h at 4°C with gentle rocking. In parallel, immunoprecipitations (IPs) with an unrelated antibody (anti-urokinase plasminogen activator receptor) or no antibodies (mock) were also performed as control. Beads were washed five times with wash buffer (0.1% SDS, 1% triton, 150 mM NaCl, 20 mM Tris-HCl pH 8.1, 1 mM PMSF) and three times with final wash buffer (0.1% SDS, 1% triton, 2 mM EDTA, 500 mM NaCl, 20 mM Tris-HCl pH 8.1, 1 mM PMSF). Bound chromatin was eluted with 1% SDS and 0.5 μg/μl proteinase K for 3 h at 50°C. DNA was de-crosslinked overnight at 65°C and purified by phenol/chloroform extraction.
Bacterial artificial chromosome microarray preparation and hybridisation
The bacterial artificial chromosomes (BACs) listed in supplemental table S1 were spotted on coverslips with the Spectral Genomics spotter (www.spectralgenomics.com). Each spot was replicated eight times. Salmon sperm and Drosophila DNA were spotted as negative controls. Male and female genomic DNA were positive controls. The array also contained control BACs from chromosome 16p13, previously shown as an open chromatin domain.17 BACs were random amplified as described (www.sanger.ac.uk) with primers designed to avoid amplification of bacterial DNA. To validate the amplification of the BAC clones, we amplified 19 random sequence tag sites (STS). All the polymerase chain reactions (PCRs) successfully amplified DNA of the expected size from the expected BAC clones (not shown).
Chromatin was amplified as described by the Farnham laboratory (http://www.genomecenter.ucdavis.edu/farnham/protocol.html) and products were run on agarose gel for quality assessment. Total genomic DNA was digested with EcoRI. Probe DNA was labelled with cyanine3 (Cy3) or cyanine5 (Cy5) dye (BioPrime DNA labeling System, Invitrogen, Carlsbad, California, USA). 0–5–1 μg of the two DNAs to compare, labelled in the two opposite ways, were co hybridised with 50 μg of human COT1 DNA in 45 μl of hybridisation buffer in a hybridisation chamber (Corning, New York, USA) at 42°C overnight. Coverslips were scanned and analysed with the Genechip software and data were normalised with local background option.18 Ratios were calculated between the hybridisation values obtained for IP and for input DNA, labelled with both dye combinations. As a quality control, the array was first hybridised to male and female DNAs (supplemental figure S2 available online).
Real-time PCR and data analysis
IP DNA was analysed by real-time PCR with Syber Green Universal Mix in a Light Cycler (Roche Diagnostic, Basel, Switzerland). The PCR primers are listed in supplemental table S3. Total DNA, IP DNA and mock IP DNA were quantified using picogreen (Molecular Probes, Carlsbad, California, USA) and equal amounts of DNA (0.5 ng) were used in PCR. Samples were run in duplicate and each experiment was repeated two to three times, on independent chromatin preparations. Data quantification was done as described by Litt et al.19 In brief, the Ct (threshold cycle) was determined and the fold increase was calculated as 2(inputCt-IPCt). All the values were corrected for non-specific signal, by subtracting the values of mock IP.
Reverse transcriptase (RT) real-time PCR was conducted as described9; p values were calculated as two tailed Student t test.
Bioinformatics analysis
Gene expression data were obtained from the GNF Symatlas database v1.2.4 (http://symatlas.gnf.org/SymAtlas/): data was from Mouse geneAtlas GNF1M, gcRMA. AD20 was used as a measure of the expression. Only genes presenting average difference (AD) ⩾200 (as described in Su et al20 at least in one of the tissues analysed) were considered in the calculation.
The number of genes and of CpG islands were downloaded from NCBI (human build 36.2, mouse build 37.1, http://www.ncbi.nlm.nih.gov/mapview). LINEs, SINEs and GC content were downloaded from the UCSC genome browser (mm8 and hg18, http://genome.ucsc.edu).
Global histone methylation data from Barski et al21 were downloaded from the summary windows, displaying the number of tags in 200 bp windows, except for CTCF where 400 bp windows were considered. A cut-off >1 tag was applied to eliminate background signals.
Results
Specific downregulation of gene expression in the POF CR1 in oocytes
We previously showed that the global downregulation of the active X chromosome occurring in mammalian oocytes10 11 was due almost exclusively to the POF critical region as a whole.9 We further analysed the region in the mouse where the CR1 syntenic region, from 101–128 Mb, is contained in a large human–mouse syntenic region, from 62.8–114.4 Mb in human and from 91.3–142.8 Mb in the mouse (NCBI build 37) (figure 1). As in human, the mouse region is gene poor (figure 1A), and enriched in LINE sequences. LINEs represent 40.48% of the region compared to 19.2% in the whole genome, while SINEs are reduced to 3.4% compared to 8.2% in the whole genome.22
Bioinformatics analysis of X chromosome gene expression in different mouse tissues. (A) Number of genes/Mb calculated as described in Materials and methods. The average gene expression/Mb in (B) oocytes, (C) fertilised eggs, (D) blastocyst, (E) embryo at 6.5 days, (F) ovary, (G) hypothalamus, and (H) lung was calculated from the sum of the normalised GNF expression values for each Mb along the X chromosome, divided for the number of genes/Mb. The region corresponding to the critical region 1 (CR1) is boxed. Dotted lines indicate the human/mouse syntenic region and the critical region. Below the position in Mb the limits of the syntenic regions are indicated.
Gene expression was analysed by determining the average level of gene expression/Mb along the X chromosome. The dataset utilised was from the SymAtlas database that contains a normalised set of expression data on many different tissues (http://symatlas.gnf.org/SymAtlas/). In figure 1 (B–H) the results of the analysis of oocytes, fertilised eggs, blastocyst and 6.5 day embryos (E6.5), as well as a number of adult tissues (whole ovary, hypothalamus and lung), are shown. Gene expression in a large portion of the human–mouse syntenic region (from 85 Mb to 132 Mb), larger than CR1, appeared specifically downregulated in oocyte and fertilised eggs compared to the rest of the chromosome. It is expressed at a higher level from the blastocyst stage and in all adult tissues. This is particularly evident from the analysis of the regions immediately flanking CR1 (85–103 Mb and 124–132 Mb) that are enriched in genes, both in human and in the mouse (figure 1).
Comparative analysis of the POF CR1 and of two autosomal regions involved in POF associated balanced translocations
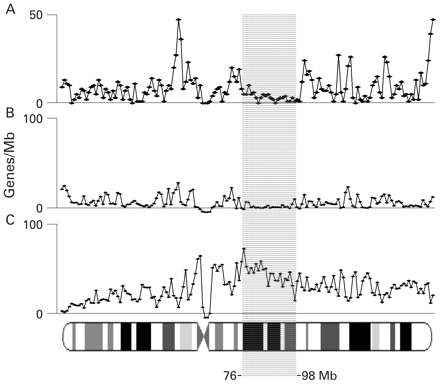
The POF CR1 spans Xq21 and overlaps with a gene poor region extending from 76–98 Mb (NCBI Build 36.1) of the human X chromosome (figure 2A). The average gene content of CR1 was 3.22 genes/Mb, about one third of that of the whole X chromosome (8.6 genes/Mb). As shown in figure 2 (B and C), the region is highly enriched in LINE sequences (46.08% of LINEs compared to the average content of the X chromosome of 31.81%), and relatively poor in SINEs (6.12% compared to 10.18% of the whole X chromosome).
Genes and repeats content of the human X chromosome, The number of (A) genes, (B) SINEs and (C) LINEs/Mb was calculated for the whole of X chromosome as described in Material and methods. The number of SINEs and LINEs is represented as the percentage of the total number of repeats. The critical region 1 (CR1) is boxed.
Within the POF CR1, a “POF breakpoint cluster” was mapped to a 1.8 Mb genomic region (from 84.4–86.2 Mb of the X chromosome) between the POF1B gene and the 3′ flanking region of the DACH2 gene.7 Compared to the surrounding area, it is enriched in LINE repeats (60.4%) and characterised by the lowest gene density (1.66 genes/Mb). Genes are large (from 102.434–684.143 Kb with an average gene length of 323.290 Kb) and cover about 50% of the sequence. In addition, the region has a low GC (36.6%) and CpG (1.4/Mb) content.
The genome organisation was compared with 1 Mb regions flanking two autosomal breakpoints in 1p35.3 and 2q14.2 described in POF patients.9 For both autosomal regions we considered 0.98 and 0.93 Mb (0.4 Mb on both sides of the breakpoints). The two autosomal regions were very different from that on the X chromosome. Both were gene dense (18 and 8 genes/Mb) containing smaller genes. Both had a high GC content (50% and 42.6%) and many CpG islands could be recognised by sequence analysis (25 and 11, respectively). As in other GC rich regions, the number of LINE repeats was reduced (15% and 37.5%) compared to the SINEs (62% and 18%).
Global analysis of histone methylation of the POF CR1
The features outlined in the previous section suggested that the POF CR1 may be an heterochromatic region. Since a large number of modifications of the N-terminal tails of histones are implicated in regulation of the chromatin state and in the establishment of global chromatin environment23 24 we analysed the chromatin organisation of CR1. We took advantage of a recently produced dataset from high resolution whole genome profiling of histone methylation.21 The data derives from Solexa IG sequencing of IP DNA from human blood cells mononucleosomes prepared using Ab to several methylated histones. To determine the level of the different histone methylations along the X chromosomes, we calculated both the number of positions/Mb where >1 DNA fragment was sequenced (grey lines in figure 3), as well as the total number of Tags/Mb, that represents the level of each chromatin modification/Mb (black lines in figure 3). As shown in figure 3A, the pattern of 2MK4H3, the histone modification associated with active chromatin showed that most of the POF CR1 (from 86 Mb to 94 Mb) was not modified by 2MK4H3. Similar results were obtained for other active chromatin methylations, such as 3MK4H3 and 1MK4H3 (not shown).
Histone methylations along the human X chromosome. The total number of sequenced Tags/Mb (black) and number of positions occupied by Tags (grey) along the human X chromosome was calculated from Barski et al21 as described in Materials and methods. Chromatin immunoprecipitation (IP) was with antibodies to (A) 2MK4H3, (B) CTCF, (D) 3MK27H3, (E) 2MK9H3 and (F) 2MK9H3. (C) Enlargement of the CTCF binding sites in the critical region 1 (CR1). The CR1 is boxed.
From the dataset, we were able to obtain data on K9H3, K27H3 and other histone methylations characteristic of silent chromatin that could be enriched in a region devoid of active chromatin modifications: the region between 88–91 Mb, corresponding to the Xq21 pseudoautosomal region (found only in human), was always devoid of any of the chromatin modifications tested. But also the surrounding portions of CR1 were free of most histone modifications studied (as an example see 3MK27H3 in figure 3D). They were, however, particularly enriched in 2MK9H3 and 3MK9H3 (figure 3E,F) in respect to the rest of the X chromosome, confirming the heterochromatic nature of the region, but also the specificity of the chromatin organisation.
Interestingly, the whole region was also low in CTCF binding sites (figure 3B). Two clusters of CTCF binding sites were mapped flanking CR1 (figure 3C): one, at 77–78 Mb, contained 209 tags; the second, at 99–100 Mb, contained 192.5 tags. The concentration of CTCF sites between the two peaks was much lower (56.85 tags/Mb) and suggested the presence of a large and homogeneous domain of heterochromatin spanning the CR1.
Similar results were obtained with the analysis of similar datasets from mouse cells (ES or MEF), on 3MK4H3, 3MK27H3 and CTCF,25 indicating that the organisation described for the human CR1 is an evolutionary conserved feature of the mammalian genomes.
Comparison of the chromatin modification of CR1 and of autosomal breakpoint regions involved in POF
The genome organisation found at the POF “breakpoint cluster” and at the autosomal breakpoint regions may underscore a different pattern of gene expression and chromatin organisation.
We have previously reported that the three genes in the CR1 “breakpoint cluster” (POF1B, CHM and DACH2) had low expression in human tissues.9 Analysis of the SymAtlas database demonstrated that it was also the same in the mouse and that most genes flanking the autosomal breakpoints were instead expressed at a relatively high level (not shown).
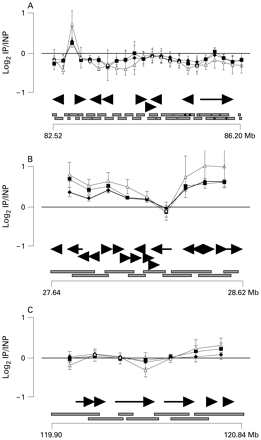
We then compared the chromatin organisation of comparable size portions of the three chromosomal regions by chromatin immunoprecipitation (ChIP) and microarray hybridisation (ChIP on chip) to look for long range chromatin modifications. We prepared a dense BAC array containing overlapping BACs (supplemental table S1 available online) corresponding to 1 Mb of chromosome 1p35.3 and chromosome 2q14.2, and about 4 Mb of the X chromosome around the POF “breakpoint cluster” (figure 4). The array was hybridised to chromatin prepared from a female lymphoblastoid cell line immunoprecipitated with Ab to the three most common histone modifications of active chromatin: 2MK4H3, AcH4 and AcH3. The silent chromatin modifications (3MK27H3, 2MK9H3 and 3MK9H3) were barely detectable in lymphoblastoid cells (not shown) and could not be studied. The hybridisation values of the IP DNA to each X chromosome BAC were consistent with the low level 2MK4H3 previously demonstrated in blood cells. Most of the normalised hybridisation values to X chromosome BACs (95.4%, 86.4% and 95.4% of the values for AcH3, AcH4 and 2MK4H3, respectively) were on the negative side of the curve in figure 4A and showed that the active chromatin modifications were few in this part of the genome. The opposite was the case for the chromosome 1 region: most of the BAC clones gave a positive hybridisation value to the three ChIP preparations (figure 4B), showing enrichment in histone acetylation and 2MK4H3. A less evident but positive enrichment could be detected on chromosome 2. In this genomic region most normalised values were only just above zero, suggesting a more limited enrichment in active chromatin modifications (figure 4C).
Chromatin immunoprecipitation (ChIP) on chip with bacterial artificial chromosome (BAC) arrays. BACs from the X (A), 1 (B) and 2 (C) chromosome regions, involved in premature ovarian failure (POF) associated translocations, were hybridised to DNAs obtained by ChIP with anti-AcH3 (black diamonds), anti-AcH4 (black squares) or anti-2MK4H3 (white triangles). The log2IP/input is shown above a representation of the gene map of each region: horizontal arrows indicate gene orientation. BACs and map positions are represented at the bottom. Only the data for one of the two dye combinations are shown: the opposite dye combination gave identical results.
The histone modifications studied mainly occurred at the promoters of active genes. To exclude the possibility that the pattern detected could be ascribed to gene or promoter density, we analysed active chromatin modifications of the putative promoters of all the genes flanking each breakpoint. The analysis was done by real-time PCR of ChIP products obtained from at least two different normal lymphoblastoid cell lines. Primers were designed in the promoter regions of each gene flanking for about 1 Mb the X chromosome breakpoint cluster (from APOOL to DACH2) and the two autosomes (from RPA2 to DNAJC8 at 1p35.3; from LOC200373 to INHBB at 2p14.2). We could demonstrate enrichment in chromatin modifications at the putative promoters of five of the seven X chromosome genes, of all six genes of chromosome 1, and six of the seven genes of chromosome 2. In all such cases two to three primer pairs within a 1 kb region were used to confirm the results. Supplemental figure S1 (available online) shows the results obtained with the LB696 cell line, used also for the global analysis. The fold increase of the ChIP over the input DNA was low for the X chromosome gene promoters (supplemental figure S1a available online) and much higher at the autosomal gene promoters (supplemental figure S1b,c available online). In agreement with the results of the global analysis, histone modifications were highly enriched at the promoters of the chromosome 1 gene and less at the promoters of the chromosome 2 genes. We can conclude that the X chromosome and the autosomal regions involved in POF associated X;A breakpoints have a different gene activity and genomic organisation, the X chromosome presenting a heterochromatic conformation and the two autosomal regions a euchromatic one.
Active chromatin modifications in POF patient lymphoblasts
As the heterochromatic state of CR1 and the euchromatic state of the autosomal regions involved in POF could be detected in lymphoblastoid cell lines, we tested whether we could also demonstrate the position effect hypothesised.9 Using the BAC array, we analysed chromatin IP preparations from lymphoblastoid cells of the two POF patients (LA1 and LB106) carrying the X;1 and X;2 rearrangements. We found no difference between patients and normal control lymphoblastoid cell lines (data not shown), suggesting that under these conditions there is no long range effect of the translocation.
In the same patients, we also analysed the putative promoters of each gene flanking the X;A breakpoints as described in the previous section, by real time PCR of the ChIP preparations. In each genomic region, we analysed one patient and four normal controls (supplemental table S2 available online). Since we had to compare chromatin modifications in cell lines presenting different global modification levels (not shown), fold increase in ChIP over input DNA was normalised by calculating the ratio between each independent modification at each point and for each cell line (the complete dataset is reported in supplemental table S2). In each genomic region, cell lines from POF patients carrying a balanced translocation in a different region were also analysed: in all instances results were identical to the normal cell lines, thus demonstrating that the POF patients’ cell lines did not carry a generalised alteration in chromatin organisation. The POF breakpoint cluster region on the X chromosome did not present any consistent difference in histone modification in any of the patients analysed (not shown). On the other hand, in patient lymphoblastoid cell lines, 2MK4H3 was altered at the promoters of some of the autosomal genes (figure 5).
Chromatin modification analysis at the promoters of autosomal genes involved in premature ovarian failure (POF) associated balanced translocations. Fold increase of chromatin immunoprecipitation (ChIP) over input DNA was calculated as described in Materials and methods and normalised by calculating the ratio between 2MK4 (black) or AcH4 (slashed) with AcH3 at each point and for each cell line. Real-time polymerase chain reaction (PCR) was done with primers in the 5′ region of the genes indicated. Data are reported in supplemental table S1. A–D correspond to the four autosomal regions: (A) 1p35.3; (B) 2q14.2; (C) 3p21; (D) 5q35. On top is the patient cell line; below is a schematic representation of each breakpoint and ChIP analysis of the two control cell lines indicated.
In 1p35.3, changes in 2MK4H3 modification could be detected at the promoter of the two genes FLJ10307 and SMPDL3B, translocated to the derX in the patient (figure 5A, supplemental table S2). In 2q14.2, 2MK4H3 was particularly increased at the 5′ end of EPB4.1L5, that was also translocated to the derX in the patient (figure 5B, supplemental table S2). In both instances the normalised values of 2MK4H3 modification for the genes indicated were either increased or decreased more than twice in patients compared to controls, while they were almost identical for the other genes in the region. This is shown in table 1 where the ratio between the normalised fold increase in patients and the average value of four normal lymphoblastoid cell controls is shown. The value for 2MK4H3 is around 0.5 (0.41–0.69) in SMPDL3B and FLJ10307 and ranges from 1.98–11.58 for EPB4.1L5. It is 1 for all the other genes, that appeared thus unchanged in patient lymphoblasts. Therefore, in patient lymphoblasts, the change in the level of 2MK4H3 appears quite relevant, considering that only one chromosome was affected.
Variation in 2MK4 in premature ovarian failure (POF) patients
To confirm the observed position effect we analysed as described above the promoters of all the genes found at two additional autosomal regions involved in X;autosome translocations. The HJ patient carried a breakpoint in 3p21, 2 Mb from the centromere, and patient LB373 in 5q35.9 None interrupted a gene, but both interrupted a gene rich region. Both promoters at 3p21 and the promoter of the MGAT1 gene in 5q35 were altered compared to controls (figure 5C,D, table 1).
No changes in histone modifications could be detected by this analysis at the 5′ end of some of the genes: a number of additional breakpoints should be analysed to explain why only some genes and possibly at one side of each breakpoint seem to be affected by the translocation as well as why the histone modification is in some cases decreased (see SMPDL3B, FLJ10307, CGGBP1 and MGC26717 in figure 5A,C) and in others it is increased (see EPB4.1L5 and MGAT1 in figure 5B,D).
A common feature of all cases studied is the limited alteration in the chromatin modification pattern in the patient as only one of the active chromatin modifications analysed appeared affected. The results suggest that transcriptional regulation in lymphoblastoid cells may not be greatly changed. In two cases (MGAT1 and SMPDL3B) we could determine by allele specific real-time RT-PCR whether the chromatin change had altered gene expression. In both cases it was not the case (not shown).
Discussion
In this paper we demonstrate an epigenetic effect of the active X chromosome on the promoters of genes translocated next to the POF CR1 and give support to our previous suggestion that POF in X;autosome balanced translocations is due to a position effect of CR1 on translocated autosomal genes.9 We also show a peculiar heterochromatic organisation of the CR1, which underscores structural and/or functional roles for this large portion of the mammalian X chromosome.
The POF CR1 is gene poor region, highly enriched in repetitive sequences and particularly in LINE retrotransposons. The heterochromatic nature of the POF CR1 was confirmed by the results on histone modifications in somatic cells as well as by gene expression analysis. The whole CR1 was devoid of most active chromatin modifications analysed, while it appeared uniquely enriched in 2MK9H3 and 3MK9H3, markers of silent heterochromatin and of centromeres.23 The organisation of the POF CR1 is also reminiscent of the centromeres for several other of its characteristics: the A/T rich base composition, the high content of LINE repeats, and the very low meiotic recombination frequency.26 We can hypothesise that, as it was shown for centromeres, RNAi encoded by the repeats in the region may be involved in nucleation of heterochromatin and eventually binding of HP1, for the maintenance of the heterochromatic state.27
We also show that the whole region was flanked by two peaks of CTCF binding sites, in human and in the mouse, and appeared instead quite devoid of CTCF binding sites. CTCF is a ubiquitously expressed and highly conserved protein, implicated in the establishment of insulators, the elements that in vertebrates prevent the spreading of heterochromatin and restrict transcriptional enhancers from activating unrelated promoters.28 29 This finding confirms the large and homogeneous chromatin domain and underscores a structural role for the region. The relative vicinity of CR1 to the XIC suggests a role for the region in this process, possibly at the X chromosome pairing and counting, one of the earliest events in X chromosome inactivation.30 31 Whether this may be related to the exceptional enrichment in LINE sequences of the CR1, as it was proposed by others,32 33 34 is an open question.
Until now, we were unable to study directly chromatin of the CR1 in oocytes or in somatic cells of the ovary; however, based on the profound downregulation of gene expression observed along CR1, up to the fertilised egg stage and to completion of meiosis, and on its low meiotic recombination frequency, it is unlikely that in oocytes CR1 may lose the heterochromatic organisation. Instead, it may acquire an oocyte specific organisation, possibly through binding of oocyte specific histone variants, that enables the transition to transcriptional quiescence and chromatin condensation and contributes to maintaining the observed inhibition of gene expression throughout meiosis. Much of the evidence and the mouse mutants suggest that this may be the case and that specific histone modifications characterise germ cell chromatin. Chromatin alterations in germ cells have been shown to produce severe consequences in progression through meiosis and oocyte maturation.13 14 35 Moreover, in the germ line, active transposition is potentially mutagenic and it is not surprising that the genomes have evolved specific mechanisms to control the activity of transposable elements in this cell type.27
When the highly heterochromatic and transcriptionally silent DNA of CR1 in oocytes is joined to euchromatic autosomal regions enriched in ovary and oocyte expressed genes, it is likely that one of the two chromatin organisations may spread into the other. In Drosophila position effect variegation, translocation next to a heterochromatic region will result in gene inactivity.36 In humans, few such examples have been described. One is the limited spreading of X chromosome inactivation to autosomes in X;A translocations.37 A second example is the long distance silencing effect of short telomeric repeats in fascioscapulohumeral muscular dystrophy.38
Analysis of the chromatin modification of the promoters of the genes flanking the breakpoints for about 1 Mb demonstrated changes in chromatin composition of autosomal gene promoters. Such changes were specific of the breakpoint regions and not found elsewhere in the patients. They confirmed the position effect. In lymphoblastoid cells, among the three chromatin modifications analysed, only one was changed and in the two cases where it could be studied the gene expression was not modified.
In conclusion, the results show that translocation to CR1 is able to modify the chromatin of autosomal genes in somatic cells, but that this is not sufficient to delete all active chromatin modifications and alter gene expression. This may, however, occur in oocytes where downregulation of gene expression is more evident and widespread, and can account for the specificity of the POF phenotype. Our results support the hypothesis of a role for CR1 in the downregulation of oocyte expressed genes during oocyte and ovarian follicle maturation and of the X linked POF as an epigenetic disorder. Analysis of the autosomal breakpoint regions in patients carrying X;A balanced translocations interrupting CR1 may therefore be a novel tool to identify candidate genes for POF.
Acknowledgments
We thank the patients for agreeing to contribute to this research and our colleagues for many helpful discussions during this work.
REFERENCES
Supplementary materials
Web only appendices 46;9:585
Footnotes
▸ Additional tables and figures are published online only at http://jmg.bmj.com/content/vol46/issue9
Funding This work was supported by MIUR-FIRB RBNE0189HM_005 and Telethon Italy.
Competing interests None declared.
Patient consent Obtained.
FR’s present address is Sbarro Institute for Cancer Research and Molecular Medicine, Temple University, Philadelphia, Pennsylavania, USA.
TP’s present address is Departments of Pediatrics and Medicine, UCSD School of Medicine, La Jolla, California, USA.
Provenance and Peer review Not commissioned; externally peer reviewed.